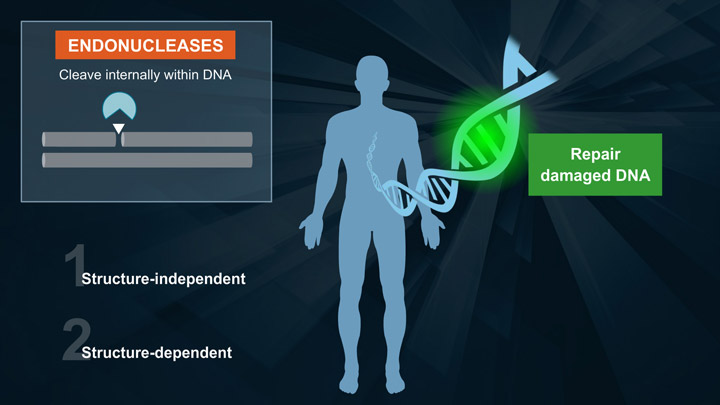
DNA Repair Enzymes and Structure-specific Endonucleases
Choose Type:
- What reaction conditions were used to define Authenticase™?
- How does Authenticase™ improve the quality and fidelity of PCR gene assembly?
- How do I convert my gene of interest into oligonucleotides?
- Why do you recommend setting up two tubes for the PCR reaction containing different amounts of Authenticase™-treated samples as templates?
- Can I use Authenticase™ for genome editing applications?
- Does Authenticase™ recognize single base pair mismatches or indels (insertions/deletions)?
- What common additives inhibit Authenticase™?
- What PCR reagents are recommended for DNA amplification in genome editing (CRISPR/Cas9, TALEN, ZFN) mismatch detection assays?
- Can I use Authenticase™ genome editing (CRISPR/Cas9, TALEN, ZFN) mismatch detection assays with unpurified PCR products?
- What size PCR amplicon should I design to analyze the genomic editing efficiency?
- If my PCR reaction yield is low, can I add more than 5 µl of the PCR reaction to the digestion reaction?
- Why do I see an extra band when I run the undigested heteroduplex on a Bioanalyzer or agarose gel?
- What are the differences between Mismatch Endonuclease I (NEB #M0678) and Authenticase (NEB #M0689)?
- Comet Assay - Modified for Detection of Oxidized Bases Using the Repair Endonucleases Fpg, hOGG1 and Endonuclease III (Nth)
- Control Reaction Protocol for PreCR Repair Mix
- Sequential Reaction Protocol for PreCR Repair Mix
- Standard E. coli DNA Gyrase Reaction (M0306)
- Standard Reaction Protocol for PreCR Repair Mix
- Using RecA and an oligonucleotide to form a stable triple helix
- Determining Genome Targeting Efficiency using T7 Endonuclease I
- T7 Endonuclease I-based Mutation Detection with the EnGen® Mutation Detection Kit (NEB #E3321)
- Transfection of Cas9 RNP (ribonucleoprotein) into adherent cells using the Lipofectamine® RNAiMAX
- Error Correction During Gene Synthesis (NEB #M0689)
- Supplemental Protocol 1: Generation of DNA fragments by PCR assembly of pooled oligos (NEB #M0689)
- Supplemental Protocol 2: Using colony PCR to identify positive clones (NEB #M0689)
- Mismatch Detection Assay (NEB #M0689)
- Transfection of EnGen® Spy Cas9 HF1 (NEB #M0667) into adherent cells using the Lipofectamine® RNAiMAX System
-
DNA Damage - the major cause of missing pieces from the DNA puzzle
- Activities of DNA Repair Enzymes and Structure-specific Endonucleases
- Properties of DNA Repair Enzymes and Structure-specific Endonucleases
- DNA Damage and PreCR
Feature Articles
Selection Tools
Usage Guidelines
- Karbaschi, Mahsa., Macip, Salvador., Mistry, Vilas., Abbas, Hussein H.K., Delinassios, George J., Evans, Mark D., Young, Antony R., Cooke, Marcus S. (2015) Rescue of cells from apoptosis increases DNA repair in UVB exposed cells: implications for the DNA damage response Toxicol Res; 4, 725-738. DOI: doiL 10.1039/c4tx00197d
- Mauris, J.and Evans, T.C., Jr. (2010) A human PMS2 homologue from Aquifex aeolicus stimulates an ATP-dependent DNA helicase. J Biol Chem; 285(15), 11087-11092. PubMedID: 20129926
- Gehring, A.M., Zatopek, K.M., Burkhart, B.W., Potapov, V., Santangelo, T.J., Gardner, A.F (2019) Biochemical reconstitution and genetic characterization of the major oxidative damage base excision DNA repair pathway in Thermococcus kodakarensis DNA Repair (Amst); PubMedID: 31841800, DOI: 10.1016/j.dnarep.2019.102767
Products and content are covered by one or more patents, trademarks and/or copyrights owned or controlled by New England Biolabs, Inc (NEB). The use of trademark symbols does not necessarily indicate that the name is trademarked in the country where it is being read; it indicates where the content was originally developed. The use of this product may require the buyer to obtain additional third-party intellectual property rights for certain applications. For more information, please email busdev@neb.com.
This product is intended for research purposes only. This product is not intended to be used for therapeutic or diagnostic purposes in humans or animals.