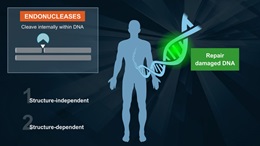
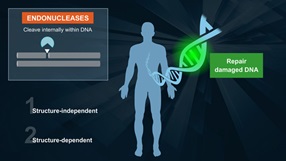
Endonuclease IV
- DNA AP endonuclease
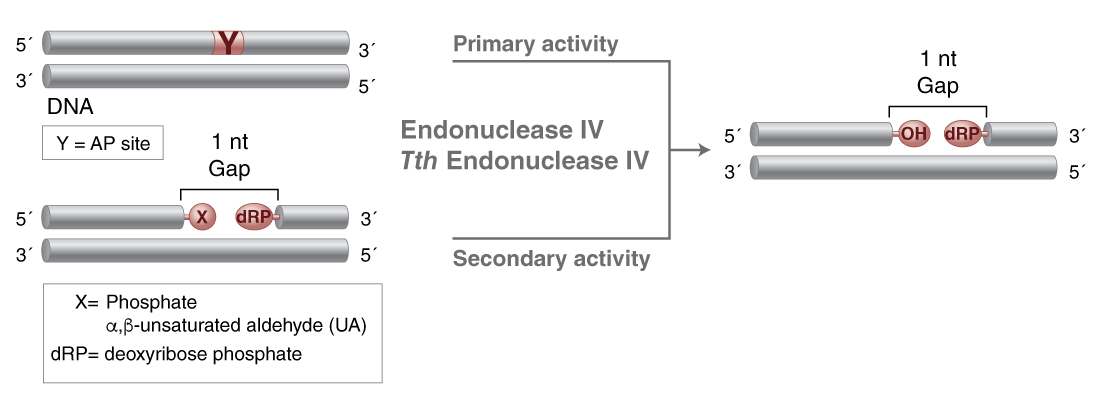
- Catalyzes the cleavage of DNA phosphodiester backbone at AP sites via hydrolysis leaving a 1 nucleotide gap with 3'-hydroxyl and 5' deoxyribose phosphate (dRP) termini
- Also has 3'-diesterase activity which can remove 3' phosphate, 3'-α, β-unsaturated aldehyde, phosphoglycoaldehyde, and other 3' blocking groups
Featured Video
-
Product Information
Endonuclease IV can act on a variety of oxidative damage in DNA (1). The enzyme is apurinic/apyrimidinic (AP) endonuclease that will hydrolyse intact AP sites in DNA. AP sites are cleaved at the first phosphodiester bond that is 5´ to the lesion leaving a hydroxyl group at the 3´ terminus and a deoxyribose 5´-phosphate at the 5´ terminus. The enzyme also has a 3´ -diesterease activity and can release phosphoglycolaldehyde, intact deoxyribose 5-phosphate and phosphate from the 3´ end of DNA.
Product Source
An E.coli strain which carries the cloned Endo IV gene.- This product is related to the following categories:
- DNA Repair Enzymes and Structure-specific Endonucleases Products
-
Protocols, Manuals & Usage
-
Tools & Resources
-
FAQs & Troubleshooting
-
Citations & Technical Literature
-
Quality, Safety & Legal
Featured Videos
Other Products You May Be Interested In
Ineligible item added to cart
Based on your Freezer Program type, you are trying to add a product to your cart that is either not allowed or not allowed with the existing contents of your cart. Please review and update your order accordingly If you have any questions, please contact Customer Service at freezers@neb.com or 1-800-632-5227 x 8.