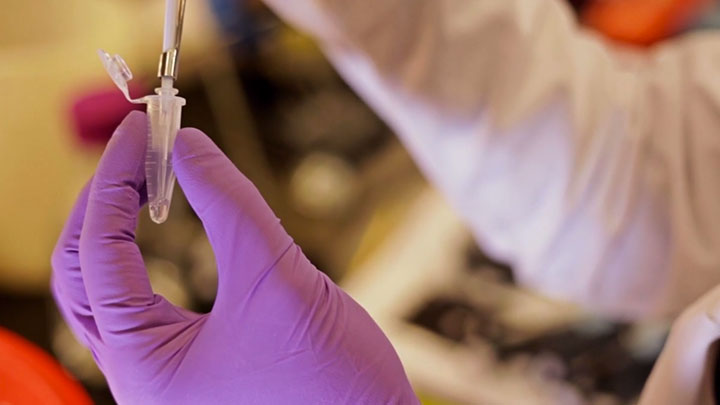
Optimizing Restriction Endonuclease Reactions
Protocols.io also provides an interactive version of this protocol where you can discover and share optimizations with the research community.
There are several key factors to consider when setting up a restriction endonuclease digest. Using the proper amounts of DNA, enzyme and buffer components in the correct reaction volume will allow you to achieve optimal digestion. By definition, 1 unit of restriction enzyme will completely digest 1 μg of substrate DNA in a 50 μl reaction in 60 minutes. This enzyme : DNA : reaction volume ratio can be used as a guide when designing reactions. However, most researchers follow the "typical" reaction conditions listed, where a 5–10 fold overdigestion is recommended to overcome variability in DNA source, quantity and purity. NEB offers the following tips to help you to achieve maximal success in your restriction endonuclease reactions.
A "Typical" Restriction Digest
| Restriction Enzyme | 10 units is sufficient, generally 1 µl is used |
| DNA | 1 µg |
| 10X NEBuffer | 5 µl (1X) |
| Total Reaction Volume | 50 µl |
| Incubation Time | 1 hour* |
| Incubation Temperature | Enzyme dependent |
* Can be decreased to 5-15 minutes by using a Time-Saver™ Qualified enzyme.
Enzyme
For additional information, please visit Restriction Enzyme Tips
- Keep on ice when not in the freezer
- Should be the last component added to reaction
- Mix components by pipetting the reaction mixture up and down, or by "flicking" the reaction tube. Follow with a quick ("touch") spin-down in a microcentrifuge. Do not vortex the reaction.
- In general, we recommend 5–10 units of enzyme per µg DNA, and 10–20 units for genomic DNA in a 1 hour digest.
- NEB has introduced a line of High-Fidelity (HF®) enzymes that provide added flexibility to reaction setup.
- Some restriction enzymes require more than one recognition site to cleave efficiently. These are designated with the “multi-site” icon
 . Please review recommendations on working with these enzymes.
. Please review recommendations on working with these enzymes.
DNA
- Should be free of contaminants such as phenol, chloroform, alcohol, EDTA, detergents or excessive salts. Extra wash steps during purification are recommended.
- Methylation of DNA can inhibit digestion with certain enzymes. For more information about methylation, Effect of CpG Methylation on Restriction Enzyme Cleavage and Dam and Dcm Methylases of E.coli
Buffer
- Use at a 1X concentration
- Supplement with SAM (S-Adenosyl methionine) to the recommended concentration if required.
Reaction Volume
- A 50 µl reaction volume is recommended for digestion of 1 µg of substrate
- Enzyme volume should not exceed 10% of the total reaction volume to prevent star activity due to excess glycerol
- Additives in the restriction enzyme storage buffer (e.g., glycerol, salt) as well as contaminants found in the substrate solution (e.g., salt, EDTA, or alcohol) can be problematic in smaller reaction volumes. The following guidelines can be used for techniques that require smaller reaction volumes.
* Restriction Enzymes can be diluted using the recommended diluent buffer when smaller amounts are needed.Restriction Enzyme* DNA 10X NEBuffer 10 µl rxn** 1 unit 0.1 µg 1 µl 25 µl rxn 5 units 0.5 µg 2.5 µl 50 µl rxn 10 units 1 µg 5 µl
** 10 µl rxns should not be incubated for longer than 1 hour to avoid evaporation.
Incubation Time
- Incubation time is typically 1 hour
- Can often be decreased by using an excess of enzyme, or by using one of our Time-Saver Qualified enzymes.
- It is possible, with many enzymes, to use fewer units and digest for up to 16 hours. For more information, visit Extended Digests with Restriction Endonucleases.
Stopping a Reaction
If no further manipulation of DNA is required:
- Terminate with a stop solution (10 µl per 50 µl rxn) [1x: 2.5% Ficoll®-400, 10mM EDTA, 3.3mM Tris-Hcl, 0.08% SDS, 0.02% Dye 1, 0.001% Dye 2, pH 8.0@25°C] (e.g., NEB #B7024 )
When further manipulation of DNA is required:
- Heat inactivation can be used
- Remove enzyme by using a spin column (NEB #T1030) or phenol/chloroform extraction
Storage
- Storage at -20°C is recommended for most restriction enzymes. For a few enzymes, storage at -80°C is recommended for periods longer than 30 days. Please refer to the enzyme's technical data sheet or catalog entry for storage information.
- 10X NEBuffers should also be stored at -20°C
Stability
- All enzymes are assayed for activity every 4 months. The expiration date is found on the label.
- Exposure to temperatures above -20°C should be minimized whenever possible
Control Reactions
If you are having difficulty cleaving your DNA substrate, we recommend the following control reactions:
- Control DNA (DNA with multiple known sites for the enzyme, e.g. lambda or adenovirus-2 DNA) with restriction enzyme to test enzyme viability
- If the control DNA is cleaved and the experimental DNA resists cleavage, the two DNAs can be mixed to determine if an inhibitor is present in the experimental sample. If an inhibitor (often salt, EDTA or phenol) is present, the control DNA will not cut after mixing.


