HpaII Methyltransferase
We are excited to announce that all reaction buffers are now BSA-free. NEB began switching our BSA-containing reaction buffers in April 2021 to buffers containing Recombinant Albumin (rAlbumin) for restriction enzymes and some DNA modifying enzymes. Find more details at www.neb.com/BSA-free.
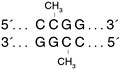
HpaII Methyltransferase recognizes the same sequence as the MspI Methyltransferase, but modifies the internal cytosine residue (C5) of the sequence CCGG.
- Supplied with 10x rCutSmart™ Buffer and 32 mM S-adenosylmethionine (SAM)

-
Product Information
Product Source
An E. coli strain that carries the cloned HpaII modification gene from Haemophilus parainfluenzae (ATCC 49669).- This product is related to the following categories:
- Methyltransferases for Epigenetics Products,
- Methyltransferases Products
-
Protocols, Manuals & Usage
-
FAQs & Troubleshooting
-
Citations & Technical Literature
-
Quality, Safety & Legal
Other Products You May Be Interested In
Ineligible item added to cart
Based on your Freezer Program type, you are trying to add a product to your cart that is either not allowed or not allowed with the existing contents of your cart. Please review and update your order accordingly If you have any questions, please contact Customer Service at freezers@neb.com or 1-800-632-5227 x 8.