E. coli DNA Ligase
E.coli DNA Ligase has been reformulated with Recombinant Albumin (rAlbumin) beginning with Lot #10232139.
E. coli DNA Ligase will ligate these substrates:
dsDNA
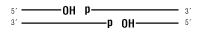
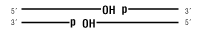
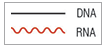
Nicked DNA/RNA
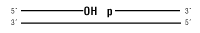
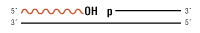
- Selective ligation of nicks in dsDNA without significant joining of dsDNA fragments regardless of end type
- cDNA synthesis
- Not sure which ligase to choose? Refer to our DNA and RNA Ligase Properties Chart or DNA Ligase Selection Chart
Featured Video
-
Product Information
E. coli DNA Ligase catalyzes the formation of a phosphodiester bond between the 5´-phosphate and the 3´-hydroxyl of two adjacent DNA strands in duplex DNA with cohesive ends. It is not appreciably active on blunt-ended substrates. E. coli DNA Ligase uses NAD as a cofactor and can be heat-inactivated. E. coli DNA Ligase is active at a range of temperatures (4° C – 37° C). Highlights
- Specific Activity: 6,000 units/mg
- Supplied with 10X Reaction Buffer
Product Source
Purified from E. coli strain containing a cloned E. coli DNA Ligase gene.- This product is related to the following categories:
- DNA Ligases Products
- This product can be used in the following applications:
- Non-Cloning Ligation
-
Protocols, Manuals & Usage
-
Tools & Resources
-
FAQs & Troubleshooting
-
Citations & Technical Literature
-
Quality, Safety & Legal
Featured Videos
Other Products You May Be Interested In
Ineligible item added to cart
Based on your Freezer Program type, you are trying to add a product to your cart that is either not allowed or not allowed with the existing contents of your cart. Please review and update your order accordingly If you have any questions, please contact Customer Service at freezers@neb.com or 1-800-632-5227 x 8.