Seamless Cloning
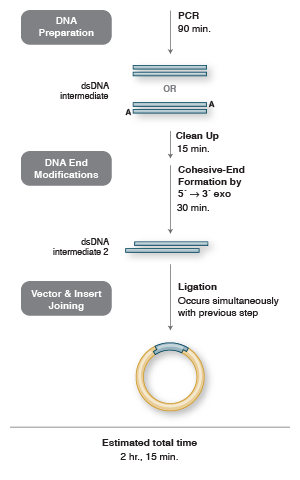
Return to Cloning & Synthetic BiologyOne seamless cloning method is the Gibson Assembly method, originally described by Daniel G. Gibson, of the J. Craig Venter Institute. His exonuclease-based method is performed under isothermal conditions after linear insert and vector are prepared by PCR and/or restriction digestion. Three enzymatic activities are employed: a 5’ exonuclease generates terminal cohesive ends (overhangs), a polymerase fills in the gaps of the annealed single-stranded regions, and a DNA ligase seals the nicks.
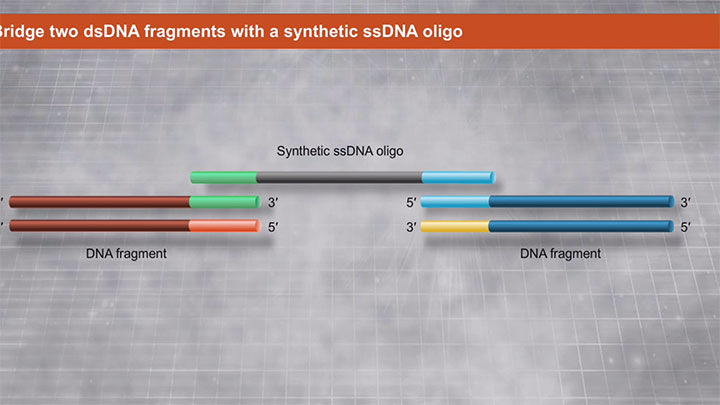
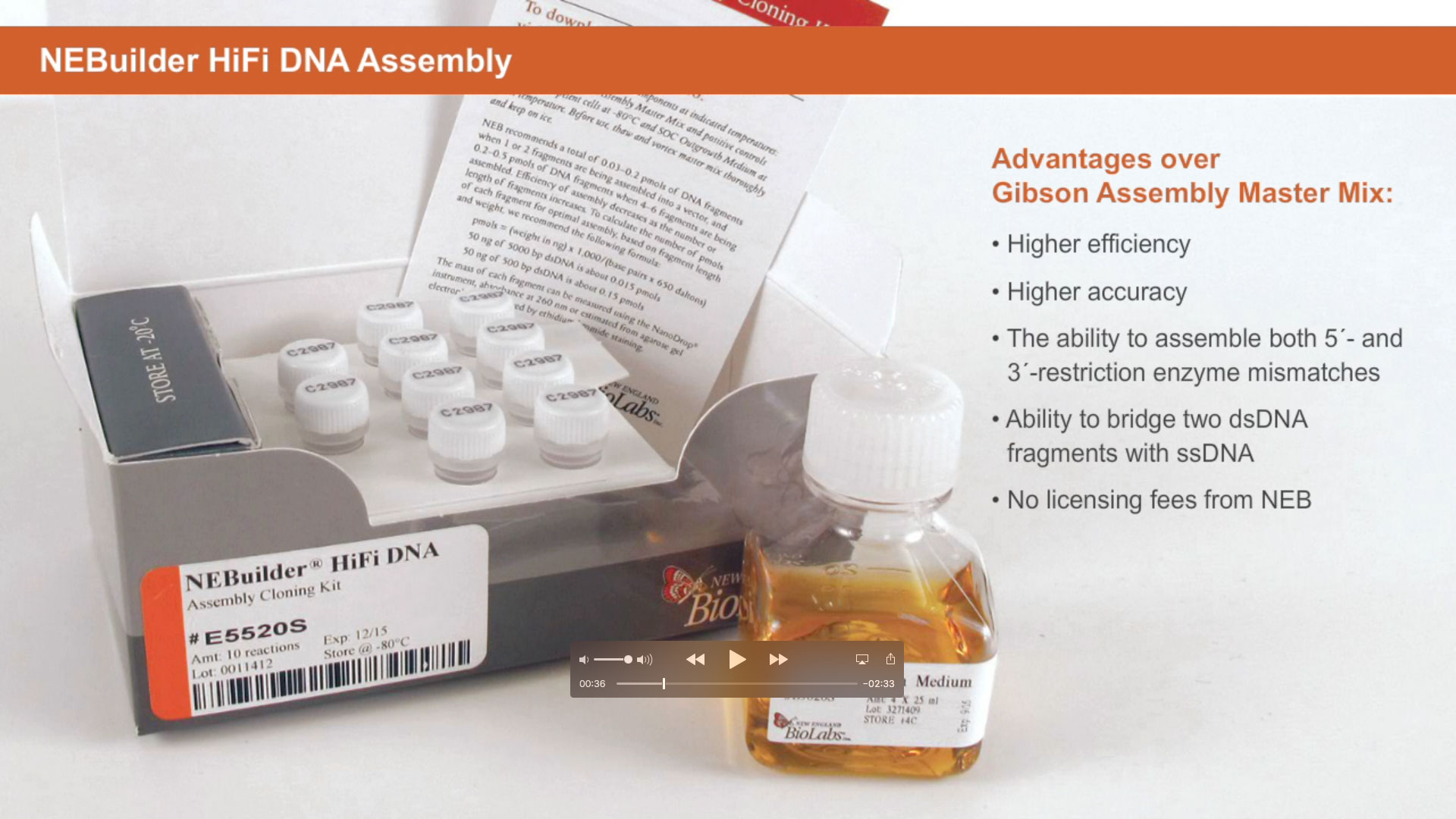
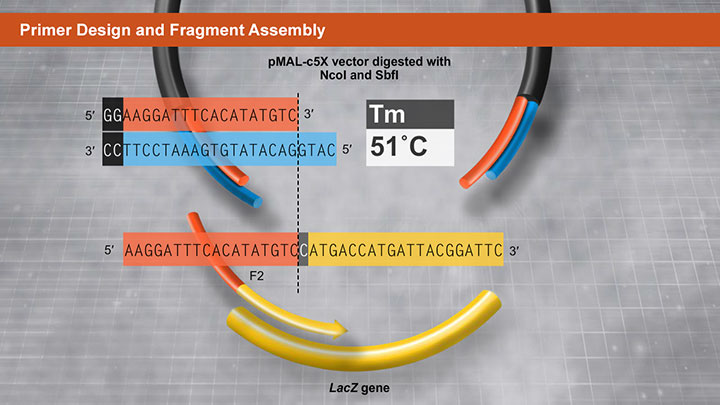
Another method utilizing a proprietary high fidelity polymerase is NEBuilder HiFi DNA Assembly. This method enables virtually error-free joining of DNA fragments, even those with 5´ and 3´ restriction enzyme mismatches. This method has also been used to assemble either single-stranded oligonucleotides or different sizes of DNA fragments with varied overlaps (15–80 bp).
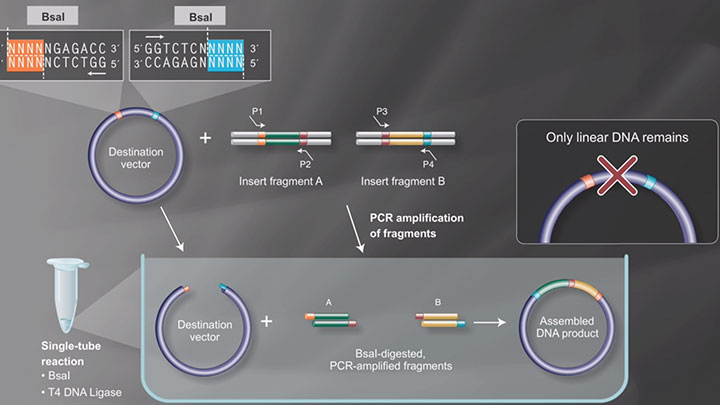
Golden Gate Assembly is another method of seamless cloning that exploits the ability of Type IIS restriction enzymes (such as BsaI-HF®v2) to cleave DNA outside of the recognition sequence. The inserts and cloning vectors are designed to place the Type IIS recognition site distal to the cleavage site, such that the Type IIS restriction enzyme can remove the recognition sequence from the assembly. The advantages of such an arrangement are three-fold:
- The overhang sequence created is not dictated by the restriction enzyme and, therefore, no scar sequence is introduced,
- The fragment-specific sequence of the overhangs allows for orderly assembly of multiple fragments simultaneously,
- The restriction site is eliminated from the ligated product, so digestion and ligation can be carried out simultaneously. The net result is the ordered and seamless assembly of DNA fragments in one reaction.
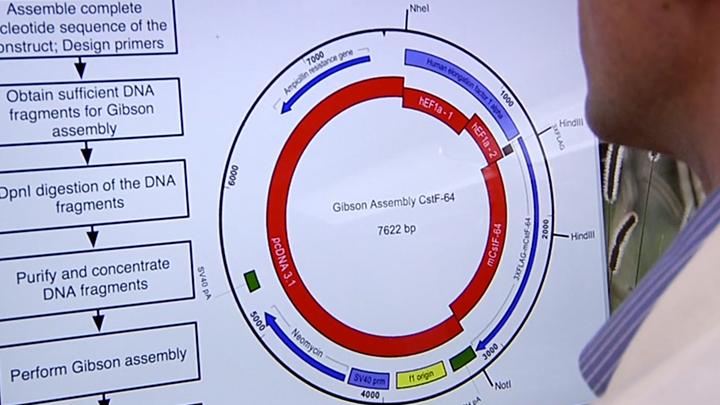
Seamless Cloning Workflow