Generation of Plasmid Vectors Expressing FLAG-tagged Proteins Under the Regulation of Human Elongation Factor-1α Promoter

Script
The overall goal of this procedure is to use the Gibson Assembly approach to create large complex DNA constructs that combine short and long segments from several, different DNA sources. First, computationally design the complete nucleotide segment from the plasmids, and then design overlapping primer sets. Next, DNA templates for the assembly reaction are amplified by PCR. Next, a purification of the PCR products is performed.
The final step is the assembly reaction transformation of the resulting plasmids and plasmid clone verification. Ultimately, the Gibson Assembly reaction is used to develop in parallel plasmids that contain flagged, tagged versions of the wild type and mutant CstF-64 proteins under the transcriptional regulation of the human elongation factor 1 alpha promoter.
Petar Grozdanov:
The main advantage of this technique over existing methods like traditional DNA cloning is that it can create in parallel similar in design wild type and mutant plasmids without sequential cloning steps.
Narrator :
Design a continuous nucleotide sequence to represent the final plasmid. Now, list the actual plasmids and DNA fragments that will be used as templates in PCR. Order DNA fragments that are not readily available, such as different combinations of tags and promoters as single or multiple synthetic, double stranded DNA fragments. Next, divide the continuous nucleotide sequence of the final construct into DNA fragments suitable for PCR. Confirm that the fragments match available plasmids and synthetic DNA fragments. Avoid DNA fragments smaller than 200 nucleotides.
Next, to access the primer generation tool, select the Set Preferences menu. In the Change Gibson Assembly Settings popup window, select the Change Prefs tab. Then, select the Build Construct menu to insert the split DNA fragments in the primer generation tool sequentially from the 5-prime to the 3-prime end. Open the Enter Vector or Insert Fragment window, paste the first DNA fragment representing the 5-prime end of the vector DNA in FASTA format, and name this DNA fragment.
Choose the appropriate way to obtain the DNA fragment, then click on the Continue tab. If there is a need to add extra nucleotides or restriction sites at the junction of the final construct, then open the Add An Insert fragment to the assembly window. In the forward or reverse primer spacer areas, enter the extra nucleotides or restriction sites to only one of the DNA fragments. Then, click on the Done tab. Repeat for all of the fragments until the construct is complete. Select the View Primers menu and review the primer sequences; also, repeat for all constructs, vector backbones and inserts, or for DNA fragments that are different.
Dilute the PCR primers to 10 micromolar in water or TE buffer. Then, dilute all the DNA templates in single stranded synthetic DNAs for the PCRs to one nanogram per microliter in water. Assemble the PCR reactions at room temperature by combining 2.5 microliters of 10 micromolar of each primer, one microliter of the one nanogram per microliter template DNA fragment, 25 microliters of hot-start proofreading DNA preliminaries, and 19 microliters of water. Mix the tube by gentle flicking and collect the liquid droplets by brief centrifugation.
Amplify simultaneously in separate tubes the DNA fragments of similar size according to the recommendations for the DNA preliminaries used. Perform 25 to 28 PCR cycles or determine the number of cycles that produce sufficient DNA yield. Resolve 5 microliters of the PCR using standard agarose gel electrophoresis. Verify that a single DNA band representing the PCR product is visible. Determine the size and relative amount of the DNA fragments using the DNA molecular weight standards. If necessary, repeat the PCR to obtain a sufficient amount of DNA fragments.
Transfer the PCRs predigested with Dpn1 to 1.5 milliliter tubes and add 81 microliters of DNA purification magnetic beads to each tube. Incubate the mixture at room temperature for 10 minutes. Place the tubes on the magnetic collector for two minutes. Now, discard the clear liquid, wash twice with 200 microliters of 80% ethanol for 30 seconds. Then, re-suspend the dry beads in 10 microliters of 10 millimolar Tris hydrochloride pH 8.0. After two minutes, spin briefly and then position the tubes on the magnetic collector for two minutes. Remove 8.5 to 10 microliters of the clear solution and place it in a new pre-labeled tube. Determine the concentration of the DNA fragments by UV spectroscopy.
Use at least 100 nanograms of DNA fragment representing the vector backbone or DNA fragment carrying the selective marker. Calculate the threefold molar excess for the DNA fragments that will be used as inserts. Mix the calculated amounts of DNA fragments in a PCR tube, then adjust the volume to 10 microliters. Add 10 microliters of the Gibson Assembly master mix. Incubate the reaction at 50 degrees Celsius in a PCR thermal cycler.
Proceed with transformation of the assembly product incompetent E. Coli or freeze the products at minus 20 degrees Celsius until needed. Follow the transformation procedure that accompanies the chemical or electrocompetent cells. Usually use two microliters of the assembly reaction per transformation reaction. Finally, confirm the DNA cloning success by DNA sequencing using specific or standard primers. Analyze the sequencing data for accuracy.
This experiment was designed to clone cleavage stimulation factor protein 64 and also mutant CstF-64 proteins fused to a 3xFLAG-tag under the expressional regulation of the HEF1-alpha promoter. A plasmid containing HEF1-alpha followed by a 3xFLAG-tag was not available to us. However, the following plasmids were available: pcDNA3.1, myc-His HEF1-alpha containing plasmid, and mouse CstF-64 plasmids. The entire sequence for the construct was assembled using the nucleotide and text editing applications.
Subsequently, the sequence was split in four convenient pieces corresponding to the available DNAs. The four DNA fragments for the assembly reactions were amplified by PCR. Resulting plasmids from individual clones were amplified and analysed by restriction enzyme digestion for proper assembly into the pcDNA vector. Select clones were then verified by sequencing.
Petar Grozdanov:
Whilst mastered, the assembly technique, including plasmid and primer design, subsequent PCR and clone verification can be performed within five days excluding the time for primer and synthetic DNA synthesis and delivery.
Related Videos
-

Introduction to Gibson Assembly® -
Introduction to NEBUILDER HIFI DNA ASSEMBLY® -
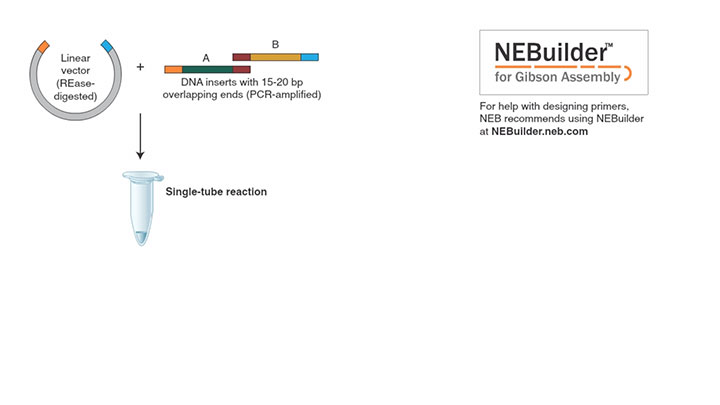

Gibson Assembly Workflow

