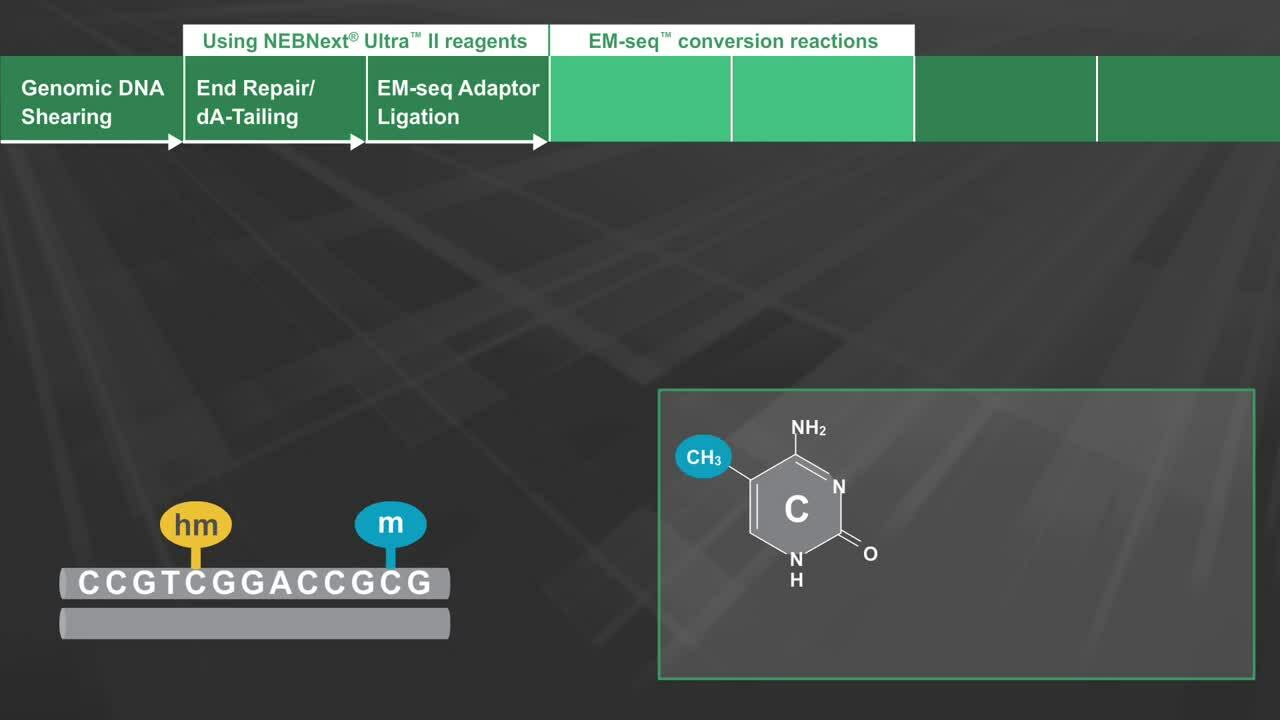
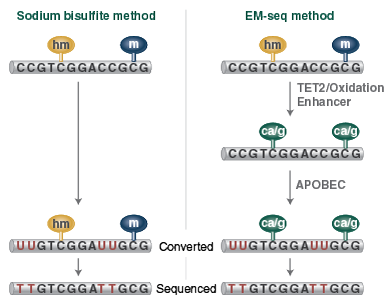
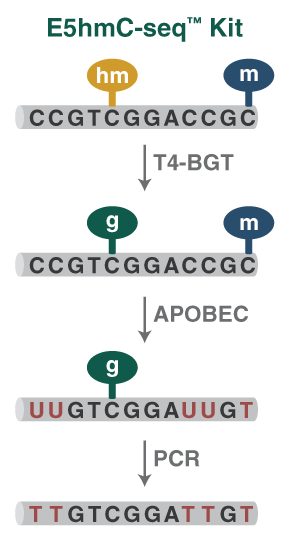
Methylome Analysis
Choose Type:
- Is it normal if the FE(II) solution from the EM-seq product is yellow or a color change is observed?
- Can the EM-seq Adaptor be substituted with another adaptor?
- Can the EM-seq Adaptor be used for bisulfite sequencing?
- What is the expected size of an EM-seq library?
- Can enzymatically fragmented DNA be used as input material for EM-seq™?
- Can EM-seq libraries be prepared from FFPE DNA?
- How should EM-seq libraries be sequenced?
- What is the concentration of the EM-seq Adaptor and EM-seq Index Primers?
- Are EM-seq libraries directional or non-directional?
- How should EM-seq sequencing data be analyzed?
- What is the concentration of the EM-seq Adaptor and Index Primers?
- What levels of conversion are typical with the control DNAs supplied in the EM-seq kit?
- Can other buffers be used in place of the supplied EM-seq Elution Buffer?
- How are EM-seq libraries prepared from cell-free DNA (cfDNA)?
- Why, at some stages of the EM-seq protocol, do the NEBNext Sample Purification Beads behave differently when cleaning up the sample?
- Are Sample Sheets available for use with the NEBNext® Multiplex Oligos for EM-seq?
- Can I use the NEBNext Ultra II FS DNA PCR-free Library Prep Kit for bisulfite conversion or EM-seq™ workflows?
- What is the difference between the NEBNext Enzymatic Methyl-seq Kit and the Enzymatic Methyl-seq Conversion Module?
- What is provided in the NEBNext Multiplex Oligos for Enzymatic Methyl-seq?
- Do I need to spike in custom sequencing primers when sequencing libraries made with NEBNext® Multiplex Oligos for Enzymatic Methyl-seq on Illumina® sequencing instruments?
- Are the NEBNext® Multiplex Oligos for Enzymatic Methyl-seq the same as NEBNext® Multiplex Oligos for Illumina®?
- What buffers are recommended for shearing DNA in NEB’s enzymatic methyl-seq (EM-seq™) workflows?
-
Enzymatic Methyl-seq: The Next Generation of Methylome Analysis
Feature Articles
Products and content are covered by one or more patents, trademarks and/or copyrights owned or controlled by New England Biolabs, Inc (NEB). The use of trademark symbols does not necessarily indicate that the name is trademarked in the country where it is being read; it indicates where the content was originally developed. The use of this product may require the buyer to obtain additional third-party intellectual property rights for certain applications. For more information, please email busdev@neb.com.
This product is intended for research purposes only. This product is not intended to be used for therapeutic or diagnostic purposes in humans or animals.