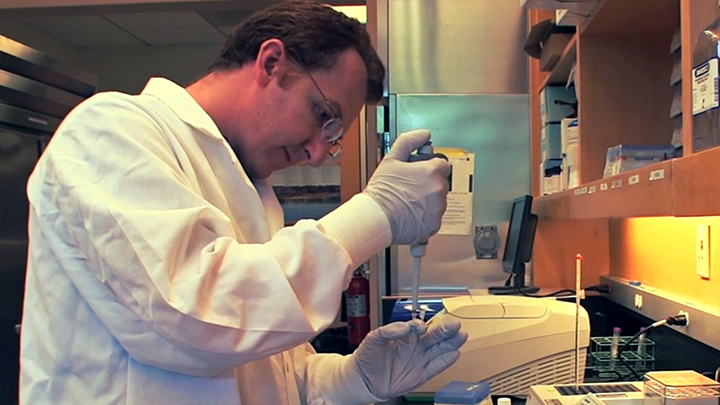
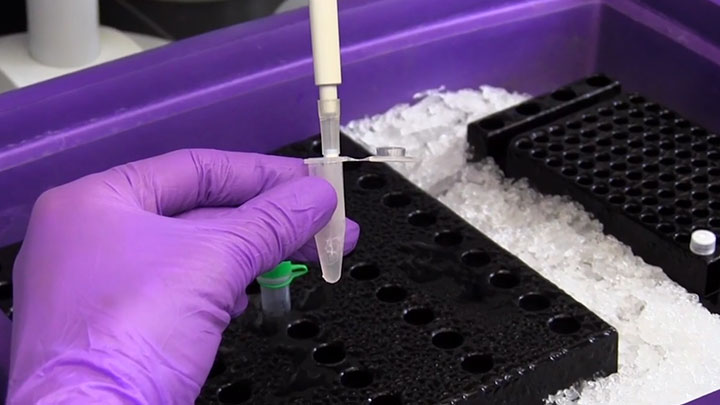
NEBNext® Ultra™ II for DNA Library Prep

NEBNext® Ultra™ II kits are designed for construction of high-quality libraries using a broad range of input amounts, from pg to µg of DNA. The workflows are fast, streamlined and automatable, with flexible product formats to enable customization.
The original Ultra II DNA Library Kits for Illumina (NEB #E7645, E7103) enable construction of libraries from pre-sheared DNA, with an input range of 500 pg to 1 µg.
NEBNext Ultra II FS DNA Library Prep – a novel enzymatic fragmentation system
We have built upon our NEBNext Ultra II DNA library prep workflow to create a fragmentation system, the NEBNext Ultra II FS DNA Library Prep Kit. The Ultra II FS kit includes a new DNA fragmentation reagent, which is also combined with end repair and dA-tailing reagents, enabling these steps to be performed in the same tube, with no clean-ups or sample loss. The same fragmentation protocol is used for any input amount (100 pg–500 ng), or GC content.
You'll be thrilled to pieces with the result – a reliable, flexible, high-quality library prep that is fast and scalable.
Highlights of the NEBNext Ultra II FS Kit
- Perform fragmentation, end repair and dA-tailing with a single enzyme mix
- Experience reliable fragmentation with a single protocol, regardless of DNA input amount or GC content
- Prepare high quality libraries from a wide range of input amounts: 100 pg–500 ng
- Generate high yields with increased reaction efficiencies and minimized sample loss
- Use with DNA in standard buffers (TE, Tris-HCl) and water
- Save time with a streamlined workflow: ~ 2.5 hours, with < 15 minutes hands-on time
- Vary incubation time to generate a wide range of insert sizes
- Available with optional SPRIselect® beads for size selection/clean-up
View or download extensive performance data in our technical notes
![]() High-yield, scalable library preparation with the NEBNext® Ultra™ II FS DNA Library Prep Kit
High-yield, scalable library preparation with the NEBNext® Ultra™ II FS DNA Library Prep Kit
![]() Data presented at AGBT by Peter Ellis, Senior Staff Scientist at the Wellcome Sanger Institute
Data presented at AGBT by Peter Ellis, Senior Staff Scientist at the Wellcome Sanger Institute
![]() Improved library preparation with the NEBNext® Ultra™ II DNA Library Prep Kit for Illumina®
Improved library preparation with the NEBNext® Ultra™ II DNA Library Prep Kit for Illumina®
![]() AGBT 2018 Wellcome Sanger Institute poster: Thurston, S. et al. (2018). A Large Genome Centre Core Pipeline Refresh.
AGBT 2018 Wellcome Sanger Institute poster: Thurston, S. et al. (2018). A Large Genome Centre Core Pipeline Refresh.
Ultra II Kit overview
| Enzymatic fragmentation included | Input amounts | Available with optional SPRIselect® beads | Compatible with PCR-free workflows | Compatible with EM-seq™ and bisulfite sequencing workflows |
|||
|---|---|---|---|---|---|---|---|
| NEBNext Ultra II FS Kits (NEB #E7805, E6177) | Yes | 100 pg – 0.5 µg | Yes (NEB #E6177) | Yes | No | ||
| NEBNext Ultra II DNA Kits (NEB #E7645, E7103) | No | 500 pg – 1 µg | Yes (NEB #E7103) | Yes | Yes | ||
What people are saying about the NEBNext Ultra II FS Kit
The Wellcome Sanger Institute currently processes thousands of DNA samples each month via its core DNA sequencing library construction pipelines. However, a recent requirement to generate high quality whole genome and targeted sequencing data from biopsy material led us to develop and implement a novel workflow enabled by NEBNext Ultra II FS. Our new automated workflow, coupled with the high efficiency of the NEBNext Ultra II FS reagent, is allowing us to routinely generate deep sequence data from as few as 100-1000 human cells.
In our core, we process large number of samples for genomics applications. Even with the high-throughput Covaris® model, the sample shearing is still a bottle neck for us. The NEBNext DNA Ultra II FS DNA kit is such a wonderful product. First of all, it increased our throughput dramatically for DNA sample shearing. The kit combines the shearing, end-repair, dA-tailing, and linker-ligation in a couple simple steps without the need of purification in between. The whole process has been reduced from two days to a few hours. Second, we love the low input option. Because the kit eliminates the need for purification for each of the intermediate steps, it reduces a lot of sample loss during the purification steps, which enables us to use much less starting materials and saves tons on purification beads. Third, we are amazed by the randomness of the enzymatic shearing. We did not observe any significant difference from mechanical shearing. We have used this kit successfully with many large projects including high-throughput genomic screening assays.
NEBNext Ultra II FS performance is exceptional. The possibility to start with higher DNA concentrations allows for less consumption of reagents for initial QC. Obtaining the desired fragment length and less hands-on time are also key factors when preparing genomic libraries and NEBNext Ultra II FS can provide that. The use of fewer PCR cycles also decreases the bias associated with the PCR step.
This kit is fantastic! I was able to create a high quality genomic library from 200 pg of DNA using the NEB Ultra II FS Kit. I had a non-model organism, limited material, GC bias and no experience with genomic library prep. The enzymatic fragmentation meant I didn’t lose any precious material in shearing and I was more than happy with the yield.
I don’t know of any other NGS library prep kit that works so well with small genomes/DNA fragments e.g. bacterial, phage genomes, plasmids, PCR products.
Available products include:
|
NEBNext Ultra II FS DNA Library Prep Kit |
|
|
NEBNext Ultra II FS DNA Library Prep with Sample Purification Beads |
|
|
NEBNext Ultra II FS DNA Module |
|
|
NEBNext Ultra II DNA Library Prep Kit for Illumina |
|
|
NEBNext Ultra II DNA Library Prep with Sample Purification Beads |