High Molecular Weight DNA Extraction
< Return to NEBMonarch.com
The Monarch® HMW DNA Extraction Kits provide a rapid and reliable process for extracting high molecular weight DNA (HMW DNA) from biological samples. These kits utilize an optimized process that combines gentle cell lysis with a tunable fragment length generation, followed by precipitation of the extracted DNA onto the surface of large glass beads. DNA size ranges from 50-250 kb for the standard protocol and into the Mb range for certain sample types when the lowest agitation speeds are used. Purified DNA is recovered in high yield with excellent purity, including nearly complete removal of RNA, and is ready for use in downstream applications including long-read sequencing.
Reasons to Choose the Monarch HMW DNA Extraction Kits
- Extremely fast, user-friendly protocols utilizing a novel glass-bead-based approach
- Reproducibly purify high molecular weight genomic DNA (HMW DNA) from various sample types
- Tune fragment length by varying agitation speeds during lysis; achieve DNA in the Megabase size range with low speeds
- Achieve outstanding results when compared to other commercially available solutions
- Excellent performance in long read sequencing
Extremely fast, user-friendly protocols utilizing a novel glass-bead-based approach
Comparison of HMW DNA Extraction Methods| Extraction Method | Approximate protocol/workflow time* | Notes** | ||
|---|---|---|---|---|
| Cells | Blood | Tissue | ||
| NEB Monarch HMW DNA Extraction | 30 minutes | 60 minutes | 90 minutes | Fast, convenient, High yield, pure, tunable DNA size |
| Circulomics® Nanobind® | > 60 minutes | 60 minutes | >2 hours | Several hands-on steps, difficult to process some samples due to high viscosity. |
| Phenol/Chloroform Extraction | > 6 hours | > 6 hours | > 6 hours | Hazardous reagents, lengthy workflow, DNA is difficult to dissolve |
| Qiagen® MagAttract® | Not compatible | ~60 minutes | Overnight + 40 minutes |
DNA is heavily sheared |
| Qiagen® Genomic Tips | ~3.5 hours | 3.5 hours | ~4 hours | Long workflow, sub-optimal yields, not very high molecular weight |
| Revolugen® FireMonkey® | ~45 minutes | >60 minutes | Not compatible | Only second eluate is used (very low recovery); DNA heated to 80°C leading to denaturing |
**Notes are based on internal testing or usage
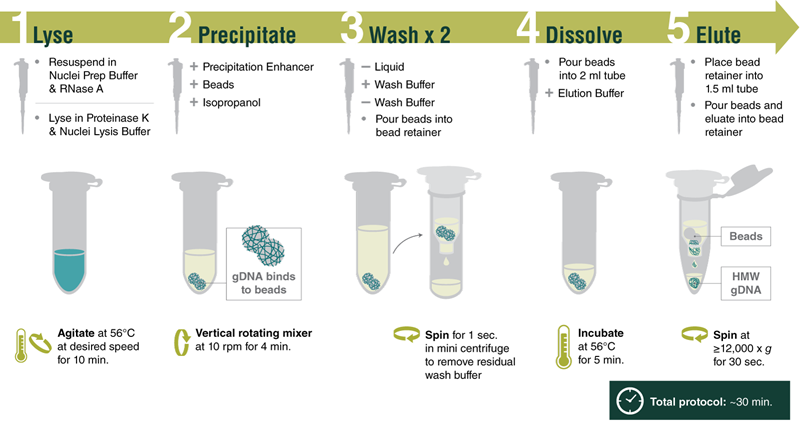
Reproducibly purify high molecular weight genomic DNA (HMW DNA) from various sample types
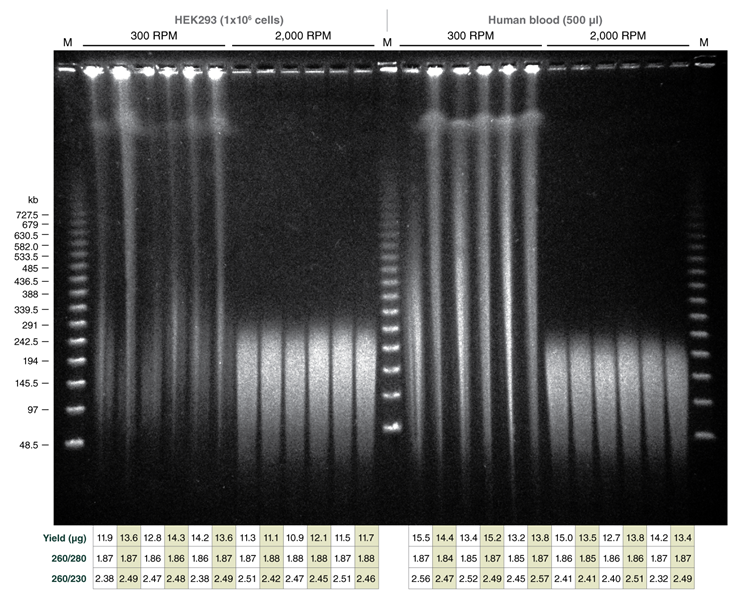
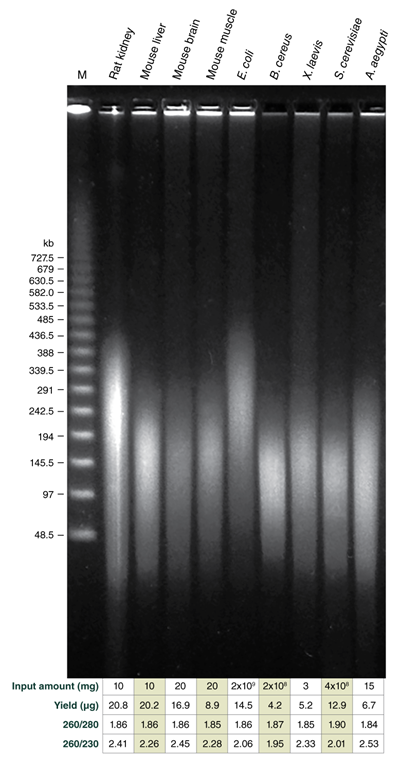
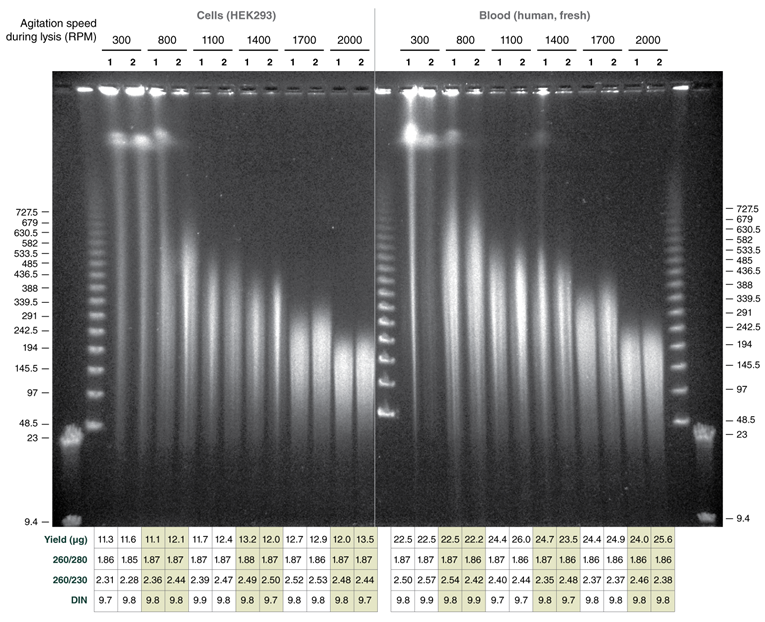
Tune fragment length by varying agitation speeds during lysis; achieve DNA in the Megabase size range with low speeds
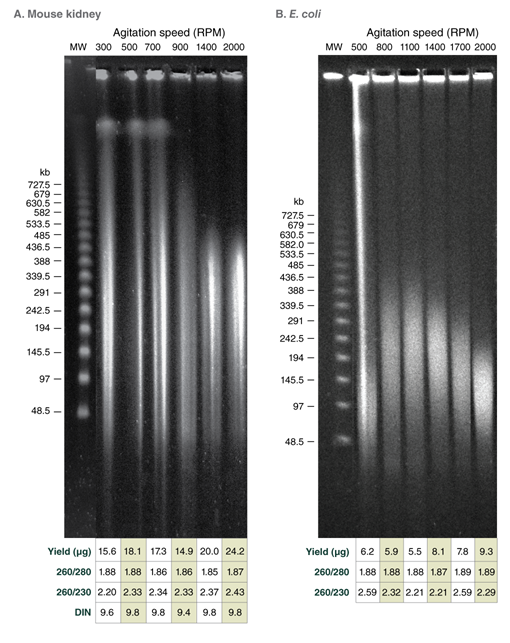


Achieve outstanding results when compared to other commercially available solutions
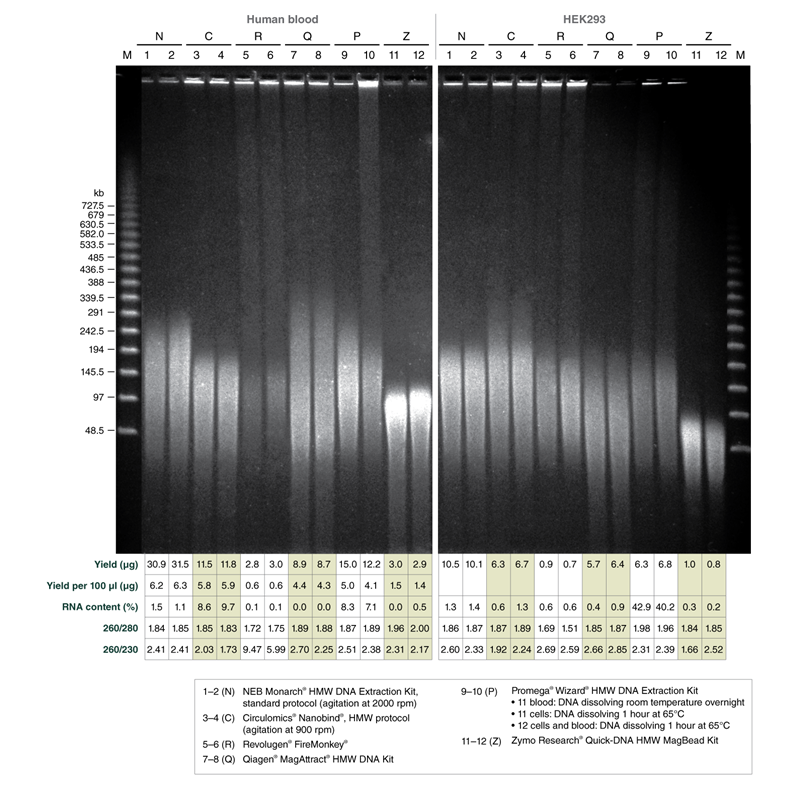
Monarch samples (lanes 1-2) were purified at maximum agitation speed during lysis (2000 rpm). Variation in fragment length of cellular DNA using the standard protocols for Monarch and Circulomics (lanes 1-2 and 3-4, respectively) results from agitation speeds during lysis (Monarch: 2000 rpm, Circulomics: 900 rpm). All other data presented are duplicate samples from each different kit and the standard protocols dedicated to blood or cells were followed. Qiagen does not provide a protocol for cultured cells; a modified version of the blood protocol was followed. Samples were eluted in 100 μl, except for Zymo which was eluted in 50 μl according to their recommendations.
Yield and purity of the standard samples were analyzed on Trinean Dropsense 16 spectrophotometer (now Unchained Labs Lunatic. Reported blood sample yields were normalized per 100 μl. RNA content was determined by HPLC analysis of nucleoside content after digestion of 1 μg of eluted DNA with the Nucleoside Digestion Mix (NEB #M0649) . The optional RNase treatment was performed with the Zymo prep.
Excellent performance in long read sequencing
| HEK293 SAMPLE 1 | HEK293 SAMPLE 2 | HUMAN BLOOD SAMPLE 1 | HUMAN BLOOD SAMPLE 2 | MOUSE KIDNEY SAMPLE | |
|---|---|---|---|---|---|
| Mean read length (bases) | 21338.9 | 19249.9 | 21522.6 | 24677.7 | 27120.7 |
| Mean read quality | 12.8 | 13.2 | 13.4 | 13.3 | 13 |
| Median read length | 10388 | 9702 | 10130 | 12593 | 23150 |
| Median read quality | 13.2 | 13.7 | 13.9 | 13.8 | 13.5 |
| Number of reads | 377687 | 633636 | 538090 | 327314 | 164000 |
| Read length N50 (bases) | 45432 | 40415 | 46542 | 51394 | 44631 |
| Total bases | 8059414490 (8.1 Gb) |
12197410796 (12.2 Gb) |
11581090785 (11.6 Gb) |
8077351338 |
4447789727 (4.4 Gb) |
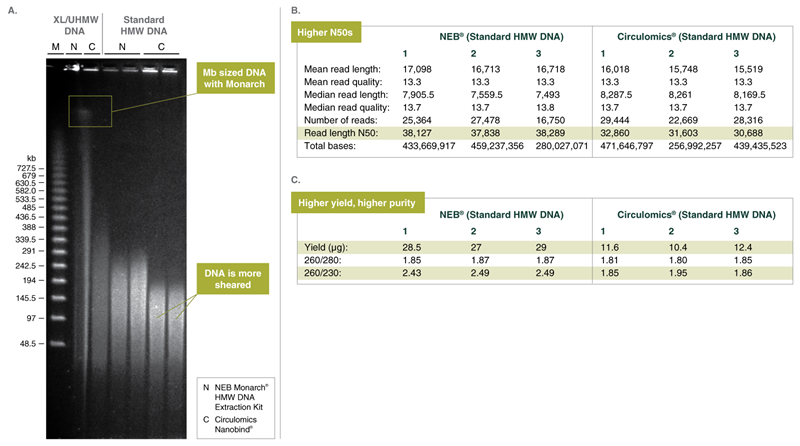
B. Oxford Nanopore Technologies ligation-based sequencing from 12-plex run with 6 blood and 6 cell samples using the Ligation sequencing kit SQK- LSK109 kit, the Native barcode expansion EXP-NBD104. FLO-MIN106D R9.4.1 flow cells and the GridION Mk1 sequencer. Results of 3 samples are shown for NEB Monarch and Circulomics. Blood samples were extracted on the same day from the same fresh human blood sample following the standard HMW protocols for blood samples from the Monarch and Circulomics kit. DNA integrity of the isolated DNA was similar to that shown in lanes 3-6 in figure A. Monarch purified samples reproducibly provided significantly higher N50 read length values.
C. Spectrophotometric analysis of yield and purity using a Trinean Dropsense 16 of the same samples shown in B. Monarch provides for significantly higher yields with the standard input volume of 500 μl versus the Circulomics workflow which is limited to 200 μl. A260/230 ratios reveal higher purity on the Monarch samples.
Featured Tools and Resources
We’ve had great success with obtaining HMW DNA for long-read sequencing from a variety of cell types, using less input and obtaining a comparable yield…It is straightforward and easy to use.