Protein Expression in E. Coli
Choose Type:
- 5 Minute Transformation (C2523)
- 5 Minute Transformation Protocol (C2527)
- 5 Minute Transformation Protocol (C2528)
- 5 Minute Transformation Protocol (C2529)
- 5 Minute Transformation Protocol (C2530)
- 5 Minute Transformation Protocol (C2566)
- 5 Minute Transformation Protocol (C3010)
- 5 Minute Transformation Protocol (C3013)
- 5 Minute Transformation Protocol (C3026)
- 5 Minute Transformation Protocol (C3028)
- 5 Minute Transformation Protocol (C3029)
- 5 Minute Transformation Protocol (C3030)
- 5 Minute Transformation Protocol (C3037)
- Expression Using SHuffle (C3026)
- Expression Using SHuffle (C3028)
- Expression Using SHuffle (C3029)
- Expression Using SHuffle (C3030)
- Expression Using NEB Express (C2523)
- Protocol for Expression Using NEB Express Iq (C3037)
- Protocol for Expression Using T7 Express (C2566)
- Protocol for Expression Using T7 Express lysY (C3010)
- Protocol for Expression Using T7 Express lysY/Iq (C3013)
- Protocol for Protein Expression Using BL21 (C2530)
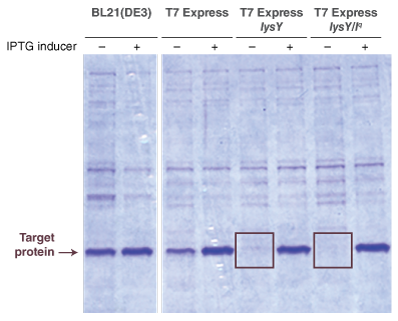
- Protein Expression with T7 Express strains
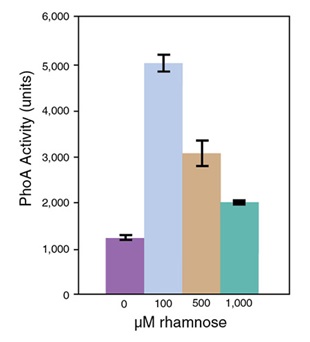
- Protein Expression Using Lemo21(DE3) (C2528)
- Protein Expression Using BL21(DE3) (C2527)
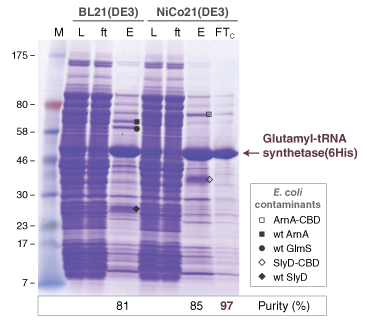
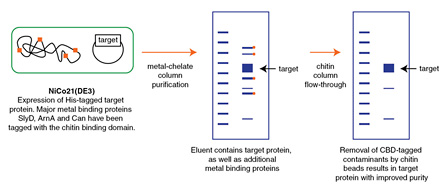
- Protein Expression Using NiCo21(DE3) (C2529)
-
Over 40 years in protein expression and purification – a historical perspective
This article provides an overview of the advances in protein expression and purification methodology over the past 40 years.
Feature Articles
- Agrawal, A., Bisharyan, Y., Papoyan, A, Bednenko, J., Cardarelli, J., Yao, M., Clark, T., Berkmen, M., Ke, N., Colussi, P. (2019) Fusion to Tetrahymena thermophila granule lattice protein 1 confers solubility to sexual stage malaria antigens in Escherichia coli. Protein Expr Purif; 153, 7-17. PubMedID: 30081196, DOI: 10.1016/j.pep.2018.08.001.
- Manta, Bruno; Berkmen, Mehmet; (2019) Disulfide Bond Formation in the Periplasm of Escherichia coli. EcoSal Plus; PubMedID: 30761987, DOI: 10.1128/ecosalplus.ESP-0012-2018.
- Leith, E.M., O'Dell, W.B., Ke, N., McClung, C., Berkmen, M., Bergonzo, C., Brinson, R.G., Kelman, Z (2019) Characterization of the internal translation initiation region in monoclonal antibodies expressed in Escherichia coli J Biol Chem; 294(48), 18046-18056.. PubMedID: 31604819, DOI: 10.1074/jbc.RA119.011008
- Reddy, P.T., Brinson, R.G., Hoopes, J.T., McClung, C., Ke, N., Kashi, L. (2018) Platform development for expression and purification of stable isotope labeled monoclonal antibodies in Escherichia coli. mAbs MAbs; 10 (7), 992-1002. PubMedID: 30060704, DOI: 10.1080/19420862.2018.1496879
- Ke, Na; Berkmen, Mehmet; Ren, Guoping; (2017) A water-soluble DsbB variant that catalyzes disulfide-bond formation in vivo Nat Chem Biol; 13, 1022-1028. PubMedID: 28628094, DOI: 10.1038/nchembio.2409
- Ren, G., Ke, N. and Berkmen, M. (2016) Use of the Shuffle Strains in Production of Proteins. Curr Protoc Protein Sci; Aug 1, 1;85:5.26.1-5.26.21.. PubMedID: 27479507 , DOI: 10.1002/cpps.11.
- Robinson, M.-P., Ke, N., Lobstein, J., Peterson, C., Szkodny, A., Mansell, T.J., Tuckey, C., Riggs, P.D., Colussi, P.A., Noren, C.J., Taron, C.H., Delisa, M.P., Berkmen, M. (2015) Efficient expression of full-length antibodies in the cytoplasm of engineered bacteria Nat Commun; (6)8072, PubMedID: 26311203, DOI: 10.1038/ncomms9072.
- Hemmis, C.W., Berkmen, M., Eser, M.and Schildbach, J.F. (2011) TrbB from conjugative plasmid F is a structurally distinct disulfide isomerase that requires DsbD for redox state maintenance. J Bacteriol; 193(18), 4588-97. PubMedID: 21742866, DOI: 10.1128/JB.00351-11
- Shouldice, S.R., Cho, S.H., Boyd, D., Heras, B., Eser, M., Beckwith, J., Riggs, P., Martin, J.L.and Berkmen, M. (2010) In vivo oxidative protein folding can be facilitated by oxidation-reduction cycling. Mol Microbiol; 75(1), 13-28. PubMedID: 19968787
E. coli Hosts
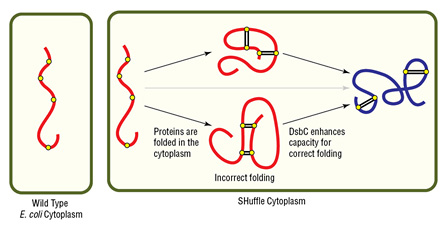
Disulfide bond formation in the cytoplasm of wild type E. coli is not favorable, while SHuffle is capable of correctly folding proteins with multiple disulfide bonds in the cytoplasm.
Products and content are covered by one or more patents, trademarks and/or copyrights owned or controlled by New England Biolabs, Inc (NEB). The use of trademark symbols does not necessarily indicate that the name is trademarked in the country where it is being read; it indicates where the content was originally developed. The use of this product may require the buyer to obtain additional third-party intellectual property rights for certain applications. For more information, please email busdev@neb.com.
This product is intended for research purposes only. This product is not intended to be used for therapeutic or diagnostic purposes in humans or animals.