Protocol for Isolation of Genomic DNA (gDNA) from up to 500 µl mammalian blood using the Monarch® Genomic DNA Purification Kit (NEB #T3010) and RBC Lysis Buffer (NEB #T3051)
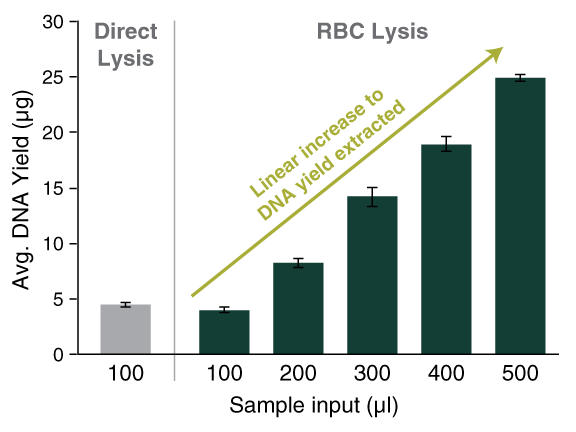
The below workflow is designed for processing higher volumes of blood (up to 500 µl) while generating similar DNA yields and concentration per 100 µl blood as obtained with the standard protocol for extraction of blood in lower volumes (up to 100 µl) – for example, if using the standard blood protocol of T3010 for gDNA isolation from a 100 µ blood sample generates 4 µg DNA, extracting gDNA from a higher 500 µl amount of the same blood sample should result in approximately 20 µg gDNA using the below protocol.
Materials Required but Not Supplied:
- Monarch® RBC Lysis Buffer (NEB #T3051) – this lysis buffer is not included as part of the Monarch Genomic DNA Purification Kit
- Monarch® Genomic DNA Purification Kit (NEB #T3010)
- One 2 ml reaction tube (or 1.5 ml tube, depends on blood volume processed) per sample
- 100 µl cold PBS per sample
Before You Begin:
- If using fresh blood, do not use blood that is older than one week (7 days) for this workflow.
- Ensure that the RBC Lysis Buffer, PBS, and centrifuge are cooled to 4°C.
- Follow recommendations from the standard protocol for extracting gDNA from 100 µl blood for storage of reagents and other specific instructions.
PART 1: SAMPLE LYSIS
- If starting with fresh blood:
- Place 150-350 µl blood in a 1.5 ml reaction tube, or 350-500 µl blood in a 2 ml reaction tube, and add 3 volumes cold RBC Lysis Buffer. For example, to 500 µl blood add 1500 µl cold RBC Lysis Buffer.
- Mix by inverting several times.
- Incubate on ice until the red blood cells have lysed. The solution will change appearance from turbid to transparent red. Depending on the species and age of the blood this will take 3 – 12 minutes, with fresh blood taking longer than older samples. Do not use blood that is older than one week (7 days) for this workflow.
- Once lysis of the red blood cells becomes apparent, leave on ice for an additional 1-2 minutes for the lysis process to complete.
- If starting with frozen blood:
- To 150-350 µl frozen blood in a 1.5 ml reaction tube, or 350-500 µl frozen blood in a 2 ml reaction tube, add 3 volumes of cold RBC Lysis Buffer to frozen sample.
- Mix by shaking briefly manually so that the liquid above the frozen blood starts to turn red.
- Place sample in an end-over-end rotator at 10-20 rpm until thawed (takes approx. 5 minutes for 500 ul blood samples).
- After sample is completely thawed vortex briefly to ensure that remaining cell aggregates are resuspended and place on ice.
- Centrifuge for 3 minutes at 1000 x g at 4°C to pellet the leukocytes.
- Carefully remove the supernatant by pipetting. Keep pellet facing down and pipet from the top side of the tube without disturbing the pellet. Remove as much of the liquid as possible.
Note: do not decant as too much supernatant will stay behind, resulting in reduced yield.
- Vortex to resuspend pellet. Add 100 µl cold PBS and vortex briefly to mix.
- Add 10 μl Proteinase K, 3 μl RNase A and 100 μl of gDNA Blood Lysis Buffer to the sample and mix by vortexing briefly. Sample will build up foam due to the relatively large amount of intact DNA.
- Incubate for 5 minutes at 56°C in a thermal mixer with agitation set at maximum speed (2000 rpm if available, 1400 rpm for older models).
PART 2: GENOMIC DNA BINDING AND ELUTION
- Add 400 µl gDNA Binding Buffer. Mix thoroughly by vortexing for 5-10 seconds at maximum vortexing speed.
- Continue with step 2 from the Blood protocol PART 2: GENOMIC DNA BINDING AND ELUTION.

