Which DNA ligase is the best choice for DNA assembly?
Posted on Thursday, July 14, 2022
By
Topic: What is Trending in Science, Tips for the lab

Katie Bilotti Ph.D. is a molecular enzymologist at NEB in the Lohman Lab, whose favorite aspect of her work is when basic discoveries can be used to improve biotechnology methods. Research on DNA ligases offers a terrific opportunity to meet this aspiration. DNA ligases are required by all organisms for replication, recombination and repair. The enzymes are also a linchpin in many molecular workflows.
In this interview, Katie explains the significance of the Lohman lab’s recent Nucleic Acids Research publication, Mismatch discrimination and sequence bias during end-joining by DNA ligases, which is the latest chapter in a several years long effort to deeply characterize end joining by DNA ligases. Excitingly, the work also shows how complex DNA assembly reactions can be strongly enhanced by the choice of ligase.
1. What is unique about ligase research at New England Biolabs?
As a part of the Molecular Enzymology research division, my goal is to have a deep understanding of enzymes. We do this by characterizing known enzymes, discovering new enzymes, and then creatively applying our knowledge of these molecular tools to biotechnology. We are lucky within the research department to be able to do basic science research as well. We're interested in DNA ligases because of their important roles in biology, including in DNA replication and repair, as well as their use in biotechnology applications. Our goal is to be the world's experts in these enzymes.
2. Why are DNA ligases important in modern biotechnology?
DNA ligases are essential enzymes to many biotechnology workflows, including traditional cloning, next generation sequencing, diagnostic assays, and DNA assembly.
3. How did your lab perform a comprehensive profiling of DNA ligases, and what are the advantages of that technique?
We use Pacific Biosciences® single molecule real time sequencing, or PacBio SMRT® sequencing for this assay. While most people use PacBio sequencing to perform long read sequencing and assemble genomes, we have used it as a tool for high throughput enzymology. We were able to evaluate the ligation of every possible 5´ four base overhang sequence in a single reaction by carefully designing a substrate oligo containing overhangs with degenerate sequence. It would have been impossible to do this study by measuring the activity on each possible overhang sequence combination in parallel using small oligos or something similar. That would involve over 65,000 different combinations of sequences! Instead, by using the PacBio sequencing-based assay that we've developed, we were able to multiplex the reaction and measure the ligation of all of those different possible sequences in just one single reaction. Additionally, because PacBio is a single molecule technique, we're able to decipher the specific mismatches for each individual ligation event that occurred, and that's something we wouldn't have been able to do if we were not looking at single molecule level.
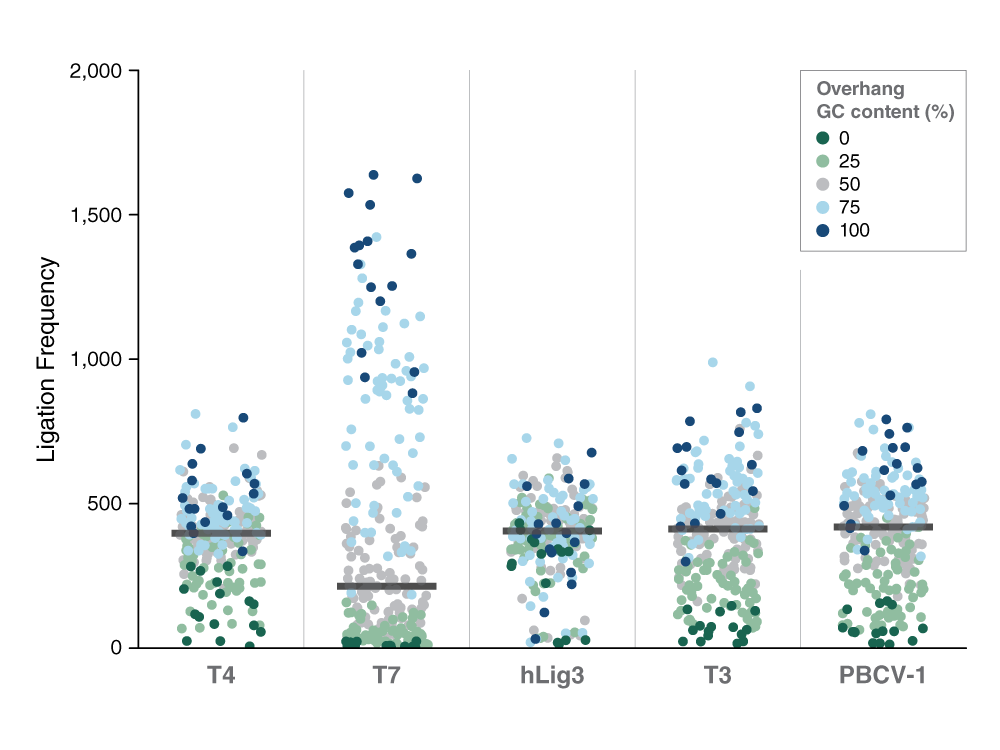
SMRT sequencing of ligation reactions with multiplex four-base overhang substrate with degenerative sequence shows how ligation sequence bias impacts efficiency. Each dot represents a different four base overhang sequence. T7 DNA Ligase is significantly more biased than other DNA ligases. (Fig.1. Bilotti et al. Nucleic Acid Research)
4. Can scientists use the findings in this study to improve ligation reactions in workflows?
Yes, this type of information can be applied and has been applied in some of our other papers. In general, this study allowed us to understand more about the basic biochemistry of ligases. We’ve characterized the enzymes in a way that contributes to the understanding of, their mechanism, their tolerance for mismatches, and their sequence preferences. The information can be used to optimize workflows by ensuring high fidelity and high efficiency ligation. More specifically, we've used this type of data for our data optimized assembly design workflows which enable high complexity, high fidelity Golden Gate Assembly. The insights have been fed into our NEBridge® Ligase Fidelity Tools that scientists can use to optimize DNA assemblies, allowing for higher complexity within a single pot reaction.
5. Is there a stellar example of that?
Yes, absolutely. We just had another paper published where we did a rapid one pot assembly of the T7 bacteriophage genome using the tools generated in this project. That's why I really like doing this kind of research, because we get to investigate the basic biochemistry of these interesting enzymes, and also apply the knowledge we gain to other scientific questions.
6. Why did your group choose to examine T3 DNA Ligase, T4 DNA Ligase, SplintR DNA Ligase, human DNA ligase 3 and T7 DNA Ligase?
We chose to examine these ligases because we wanted to look at all the ligases in the NEB catalog that are known to perform efficient end joining in the absence of accessory proteins. That makes the context a bit more relevant to biotechnology where we typically don’t want to be adding in lots of extra accessory proteins to help these enzymes do their jobs. The ligases that are part of the NEB catalog are T3, T4, T7, and the PBCV-1 DNA ligase mentioned in this study is our SplintR® Ligase. Human DNA ligase 3 (hLig3), is not sold by NEB but was an interesting example of a mammalian ligase that can catalyze end joining reactions.
7. Do you see certain sequence biases among the DNA ligases? When does GC content affect DNA ligation bias?
In this work overall, we found that different DNA ligases have different sequence preferences and biases. Ligases can have a bias that is highly depend on the GC content of the overhang sequence where overhangs with a higher GC content will ligate more efficiently just due to a stronger annealing of the overhang. This effect was particularly pronounced for some ligases, such as T7 DNA Ligase, which we found to be very biased, and very dependent on GC content. Other ligases like hLig3 appeared from this study to be very minimally dependent on GC content. We also see subtleties in the sequence preferences of these enzymes that go beyond this simple trend of GC content.
8. Is enzyme specific mismatch tolerance possible?
Yes, in addition to finding that we had differences in sequence bias and sequence preference between the ligases, we also found the ligases tolerate quite different mismatches. In general, most ligases tolerate G:T mismatches particularly well because they tend to be non-helix distorting. Those are really common mismatches across the board for ligases to tolerate. However, we did find that the degree of mismatch tolerance as well as the identity of other rare mismatches is different among ligases.
It's important to understand the mismatch tolerance of these enzymes from an applications perspective so that you can achieve high fidelity assemblies of DNA. That's what our NEBridge® Golden Gate Assembly tools help users do. You can imagine that in a complex assembly reaction where you have lots of different sequence possibilities you also have lots of potential for mismatches as you increase your number of fragments. Data from these kinds of assays helps you choose overhangs that will have minimal crosstalk and ultimately lead to a high-fidelity assembly.
9. Is mismatch tolerance what makes some ligases better suited to certain applications than others? Can researchers improve DNA assemblies with ligase choice?
In a lot of ways, yes, but there is a balance between the mismatch tolerance and sequence bias of a ligase that makes it particularly well suited or not so well suited to certain applications. The results of this study certainly inform what ligase somebody would choose for DNA assembly reactions. For example, in a complex Golden Gate Assembly reaction, you need to have lots of different fragments come together with both high efficiency and high fidelity. In order to have high efficiency ligation of a large number of different overhang sequences, you would want to use a ligase with high fidelity and low sequence bias.
This is why T4 DNA Ligase is the best enzyme for DNA assembly methods. T4 DNA Ligase has relatively low sequence bias paired with relatively high fidelity that is dominated mostly by G:T mismatches - which we now know and can avoid through careful assembly design. In contrast, while T7 DNA Ligase is extremely high fidelity, it is also extremely biased. The bias of T7 DNA Ligase limits the number of fragments that can be assembled using this enzyme, and ultimately makes it a less ideal choice for Golden Gate Assembly.
10. What kinds of future collaborations would be exciting for you?
We are continuing to evaluate ligase biochemistry in many different ways. We feel there's still a lot to learn - while we know quite a bit about nick ligation, end joining ligation is a little more mysterious at this point. We are currently using the insights gained from this work to guide structure-function and single molecule studies of ligase end-joining, which are being done in collaboration with researchers at the University of Wisconsin-Madison.
As far as applications of this type of data goes, we are interested in the assembly of large, complex targets that need to be produced rapidly with high fidelity. We have taken some initial steps toward this goal in our other recent publication, where we achieved a one-pot assembly of the 40 kb T7 bacteriophage genome from 52 parts (Pryor et al. ACS Synth. Biol. ).
In general, we are interested in research projects where we deeply characterize enzymes so that we understand everything about how they work, how to make them, how to apply them in the most effective ways, and maybe even how to make them into better enzymes. That kind of research expands our knowledge and enables us to improve all kinds of workflows.
NEB will not rent, sell or otherwise transfer your data to a third party for monetary consideration. See our Privacy Policy for details. View our Community Guidelines.
Products and content are covered by one or more patents, trademarks and/or copyrights owned or controlled by New England Biolabs, Inc (NEB). The use of trademark symbols does not necessarily indicate that the name is trademarked in the country where it is being read; it indicates where the content was originally developed. See www.neb.com/trademarks. The use of these products may require you to obtain additional third-party intellectual property rights for certain applications. For more information, please email busdev@neb.com.
Don’t miss out on our latest NEBinspired blog releases!
- Sign up to receive our e-newsletter
- Download your favorite feed reader and subscribe to our RSS feed
Be a part of NEBinspired! Submit your idea to have it featured in our blog.