
Monitoring tick and mosquito infections using LAMP: Screening for EEE and beyond
Posted on Wednesday, June 22, 2022
By
Topic: What is Trending in Science
You might say LAMP (loop-mediated isothermal amplification) was a niche application before 2020 and it took a pandemic to find its stride. Researchers involved with LAMP development always knew the potential for point-of-care and field applications — and there was a growing interest — but COVID accelerated things when RT-PCR testing infrastructure became strained. Researchers sought alternative diagnostic tools to improve accessibility. This need highlighted the utility of LAMP in settings outside a specialized molecular laboratory and brought it to the forefront. And now, field applications for LAMP for the diagnosis and surveillance of infectious diseases are gaining traction, particularly in low-resource settings.
The evolution of LAMP for diagnostics and surveillance
The Carlow lab at New England Biolabs® has used LAMP technology for infectious disease diagnostics and surveillance for almost a decade. They have developed a suite of LAMP tests to detect insect and tick-borne diseases such as Eastern Equine Encephalitis, anaplasmosis, Lyme Disease and babesiosis, and various Neglected Tropical Diseases (NTDs), such as lymphatic filariasis (elephantiasis) and onchocerciasis (River Blindness).
While LAMP does not replace PCR, the ability to amplify nucleic acids at one temperature, without the need for a thermocycler to rapidly ramp temperatures, is a huge advantage for point-of-care testing for infectious diseases; agricultural diagnostics such as testing crops for infection; and field surveillance programs involving disease vector insects and ticks. Further, colorimetric LAMP gives a visible-to-the-eye read-out, allowing scientists to dispense with specialized fluorescence detection or electrophoresis equipment to obtain results. And the ability to lyophilize LAMP reagents removes the need for refrigeration at the testing site, making it a valuable assay at low-resource locations that have no, or intermittent, electricity supply. These key features reduce the cost and make nucleic acid detection available outside the lab to populations and regions that otherwise have no access to testing.
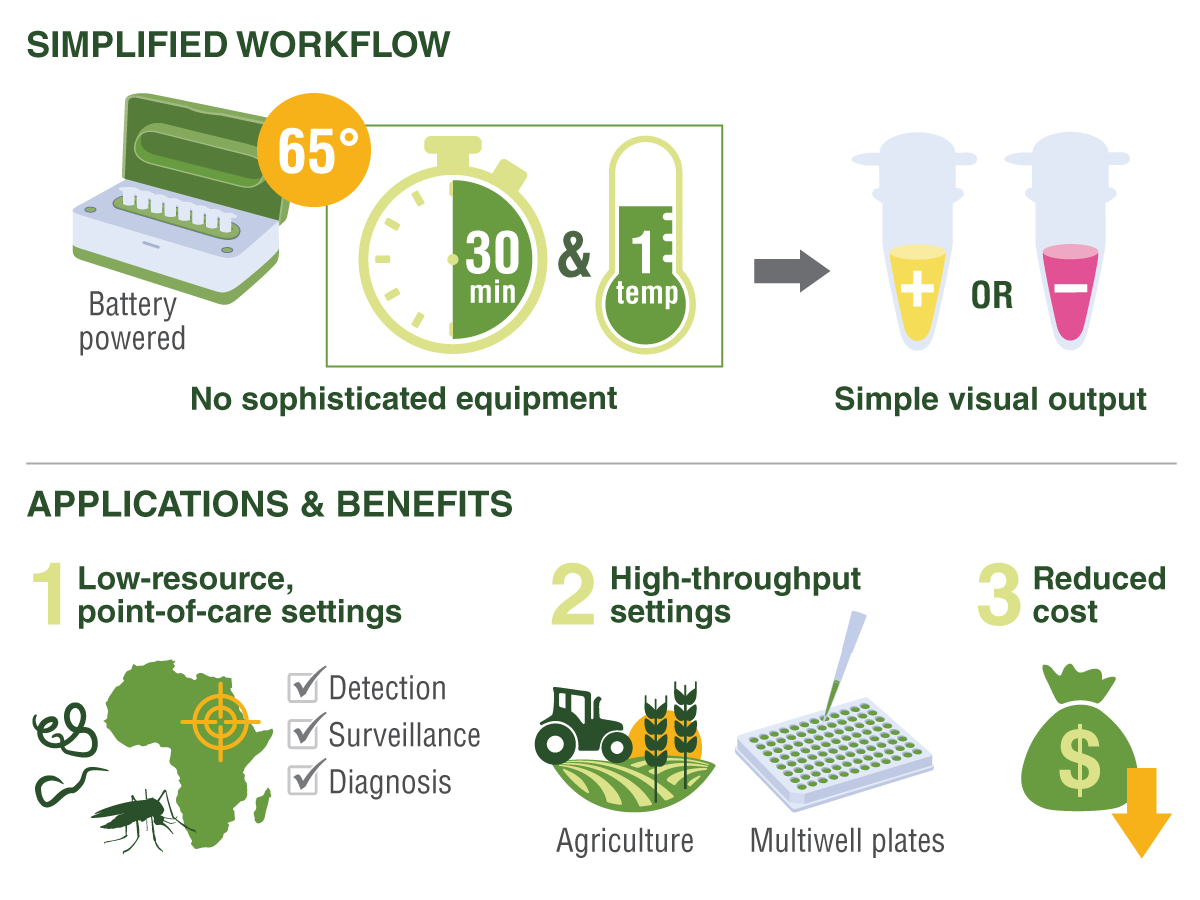
The simplified LAMP workflow requires no sophisticated equipment and can be carried out without an electrical source, with results in 30 minutes that are visible to the naked eye. LAMP is ideal for screening and diagnosing NTDs in low-resource settings, agriculture, tick and mosquito monitoring programs, and even at-home testing for infections.
LAMP uses four core and two additional “loop” primers, producing a short dumbbell structure that forms the basis for exponential amplification. The need to use a thermocycler and high temperatures to denature the DNA strands is replaced by incorporating a strand-displacing DNA polymerase, which extends the primer while displacing the duplex. You can view a more thorough explanation of LAMP in this video.
LAMP makes a LOT of DNA, and quickly. A reaction can be complete within 30 minutes. There are various outputs available for the interpretation of results. When the Carlow Lab originally started exploring LAMP, the readout was turbidity-based – it measured the precipitation of magnesium pyrophosphate in the reaction or involved a metal-based colorimetric readout using hydroxynaphthol blue. This wasn’t easy to interpret because it involves a change in color from purple to shades of blue. At this point, researchers at NEB developed another colorimetric readout of the LAMP results based on a pH change. Briefly, when the DNA polymerase adds a nucleotide, it produces a pyrophosphate and a proton. Because so much DNA is produced in a LAMP reaction, the number of protons produced is significant, and in a solution that is only moderately buffered, the pH drops by about two units. Including a pH-sensitive dye (phenol red) turns the reaction from pink to yellow, which is much easier to detect by eye.
LAMP is well-suited for diagnostics; however, it doesn’t replace PCR for some downstream applications such as cloning. This is because of the structure and length of the product, which can get very long, up to 20,000 - 25,000 bases. The products are concatemers and repeats of the LAMP amplicon sequence with branches, loops, ss/dsDNA, etc. It can be used for sequencing after fragmentation and cleanup if desired, but generally, a rapid yes/no diagnostic answer is sufficient for the intended application.
Scientists at NEB have worked to bring a suite of LAMP expertise and tools up to the molecular diagnostic standards of PCR. By 2015, they had engineered enhanced LAMP enzymes such as Bst 2.0®, controlled their activity for high-throughput and room-temperature setup with novel WarmStart® aptamers, and figured out how to prevent carryover contamination with dUTP and a new thermolabile UDG. They developed a method for LAMP multiplexing to detect multiple targets, optimized the pH-based colorimetric detection method described above, and launched master mixes and glycerol-free enzymes – an essential requirement for lyophilization.
Fast forward to 2020, and it was quickly realized that LAMP had some application in testing for SARS-CoV-2. Supply chain shortages and labs struggling to keep up with testing needs highlighted the benefit of a diagnostic test that could be applied with minimal cost, materials, and molecular expertise. This brought LAMP to the forefront as many researchers pivoted to address testing needs. NEB scientists established an in-house CLIA-certified SARS-CoV-2 LAMP assay using saliva with minimal sample prep for NEB employees.
At the same time the Carlow Lab was spearheading the CLIA lab LAMP-based testing, they were also involved in various LAMP-based collaborations, namely optimizing a colorimetric Onchocerciasis diagnostic LAMP assay in Cameroon and developing a colorimetric RT-LAMP assay for the detection of Eastern Equine Encephalitis (EEE) virus in vector mosquitoes. Read on to learn more about the latter.
Colorimetric LAMP improves the accessibility of EEEV testing in the U.S.
EEE is an arbovirus transmitted by mosquitoes. Although rare, an infection can cause brain inflammation and is often fatal. It has an enzootic cycle between mosquitoes (Culiseta melanura is the primary mosquito vector) and passerine birds. It is seasonal in some areas (such as the U.S. Northeast); however, it is endemic in Florida, providing an ecological niche for sustained wintertime circulation.
While the current gold standard for EEE surveillance is RT-PCR, it is often not implemented because of access and funding limitations. It requires expensive equipment such as a real-time thermocycler. It also requires a level of molecular lab expertise, namely the ability to run RT-PCR tests and interpret threshold (Ct) values and amplification curves. Mosquito control programs around the U.S. typically don’t have much funding, and it varies from state to state and even between counties within a state. The cost of sending samples to a specialized lab can stifle a minimally funded surveillance program.
There are 60+ mosquito control programs in Florida that monitor for multiple infections. In addition to EEE, they often monitor West Nile Virus and Saint Louis Encephalitis Virus. Some labs consist only of a cold table (to keep the mosquitoes asleep) and a field microscope (to sort them into different species morphologically). Aerial spraying is initiated once the vector populations reach a threshold level (regardless of whether or not they are infected). Another approach in many counties in Florida relies on EEE infection surveillance in chicken flocks. Samples from the chicken coops are transferred to a central arbovirus lab in Tampa, Florida, where they are assessed for seroconversion. This is a labor-intensive methodology, especially considering the work it takes to maintain chicken coops.
In a study published earlier this year (Maddison et al., 2022) in a collaboration between the Carlow lab and Tom Unnasch’s lab at the University of South Florida, a LAMP assay was developed to detect EEE virus. It was demonstrated to be a highly accurate alternative to RT-PCR. Professor Unnasch is no stranger to establishing and translating disease surveillance programs into the field; he set up the first routine diagnostic PCR testing lab for River Blindness in Africa in 1992 and works closely with the Florida Mosquito Control Association. The collaboration leveraged Carlow Lab expertise in diagnostics and primer design to produce a tremendously robust assay. The initial primer design process is critical for a successful assay, and NEB scientists and software engineers have streamlined the process with a LAMP Primer Design web tool.
The LAMP assay developed in this collaboration used three highly conserved EEE virus biomarkers (nsP3, E1, and 6K) that target different regions of the viral genome. When RT-PCR was compared with LAMP, 100% agreement was observed between the results from the quantitative (Ct) values obtained with RT-PCR and the end-point color change (pink to yellow). All three biomarkers showed a high level of sensitivity. Of the three biomarkers, E1 and nsP3 were more sensitive than 6K and comparable with the limit of detection using RT-PCR. It should be noted, however, that RT-PCR can detect down to approximately 1/10 of a virus particle, but a single infectious mosquito carries about one million virus particles. So, in that sense, all three RT-LAMP biomarkers were comparable in terms of being able to detect a single infectious mosquito.
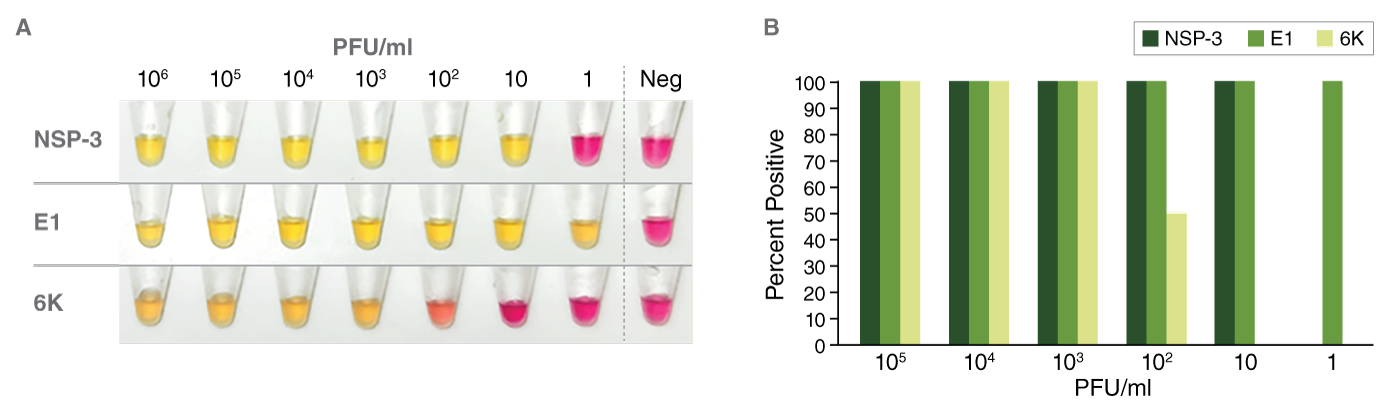
Analytical sensitivity of RT-LAMP for EEEV. (A) Colorimetric RT-LAMP assays targeting nsP3, E1, or 6K, were performed using serially diluted viral RNA (PFU/ml) as a template. A color change from pink to yellow indicates a positive result. (B) The percentage of samples that tested positive at each serial dilution (PFU/ml) of viral RNA using LAMP primers sets for nsP3 (dark green bar), E1 (medium green bar), or 6K (light green bar) are shown (calculation based on biological replicates of each PFU concentration run in duplicate with each primer set).
Additionally, pooled samples representing homogenates of more than 200 mosquitos, including Cs. melanura and also a less competent EEE virus vector, Culex erraticus, further demonstrated that RT-LAMP was a highly sensitive alternative for EEE virus detection.
Implementation of LAMP assays in local mosquito programs would require only the addition of a basic hot block (to maintain the amplification temperature at 65°C) to the existing laboratories and very little technical expertise. An added advantage is a reduction in contamination risk because of the simple color change output that can be visualized without opening tubes. The quick turnaround time to run the assay (30 minutes) enables the more extensive screening programs to increase their testing throughput.
All-in-all, this is a cost-effective, simple, fast, and accurate assay that can be implemented at the level of local testing programs; not just for EEE mosquito monitoring, but for many surveillance and diagnostic programs that would benefit from nucleic acid amplification in a simplified format.
NEB will not rent, sell, or otherwise transfer your data to a third party for monetary consideration. See our Privacy Policy for details. View our Community Guidelines.
Products and content are covered by one or more patents, trademarks and/or copyrights owned or controlled by New England Biolabs, Inc (NEB). The use of trademark symbols does not necessarily indicate that the name is trademarked in the country where it is being read; it indicates where the content was originally developed. All other trademarks are the property of their respective owners. The use of this product may require the buyer to obtain additional third-party intellectual property rights for certain applications. For more information, please email busdev@neb.com.
Don’t miss out on our latest NEBinspired blog releases!
- Sign up to receive our e-newsletter
- Download your favorite feed reader and subscribe to our RSS feed
Be a part of NEBinspired! Submit your idea to have it featured in our blog.




