Protocol for High Molecular Weight DNA (HMW DNA) Extraction from Cells (NEB #T3050)
Below is a detailed protocol for the extraction of high molecular weight DNA from cell samples. Please refer to the complete product manual for more detailed guidance. For help selecting input amounts, visit Choosing Input Amounts for the Monarch HMW DNA Extraction Kits. If you are using this kit upstream of ultra-long DNA sequencing on the Oxford Nanopores Technologies platform, please also review our Ultra-Long DNA Sequencing Workflow®’ Guidance for HMW DNA Extraction Upstream of Oxford Nanopore Technologies as well as our Protocol for UHMW DNA Cleanup in the Oxford Nanopore Technologies® UL Library Prep Workflow.
Materials Required but Not Supplied:
- Thermal mixer containing a 2 ml tube block (if not available, use a 1.5 ml block).
- Isopropanol, 275 µl per sample (Low Input: 100 µl/sample).
- Ethanol (≥ 95%).
- 1.5 ml DNase-free, low DNA binding microfuge tubes (e.g., Eppendorf® DNA LoBind®, #0030108051) are recommended for elution and storage (1 per sample); it is especially important to use low DNA binding tubes if working with UHMW DNA, which tends to bind to plastic surfaces.
- Recommended: vertical rotating mixer (e.g., Thermo Scientific® HulaMixer® Sample Mixer).
- Wide-bore pipette tips.
Important Notes Before You Begin:
- Review the complete protocol before beginning.
- Add ethanol (≥ 95%) to the gDNA Wash Buffer as indicated on the bottle label.
- Cool the Nuclei Prep Buffer to 4°C.
- Preheat thermal mixer with 2 ml block to 56°C.
- Proteinase K and RNase A should be stored at -20°C.
Starting Material Notes:
- The sample input range is 1 x 105 –1 x 107 cells.
- An input of 1 x 106 cells is recommended.
- If using > 2 x 106 cells, purified DNA will be viscous and more difficult to dissolve and handle.
- If using 300 rpm for agitation, do not exceed 5 x 106 cells.
- Use the table below to determine the designation of your sample type, which will determine various volumes in the protocol.
PROTOCOL DESIGNATION
NUMBER OF CELLS
Standard Input
> 5 x 105 – 1 x 107. 1 x 106 is recommended.
Do not exceed 5 x 106 cells if using 300 rpm agitation for “UHMW DNA”.Low Input
1 x 105 – < 5 x 105
Below 1 x 105 cells, DNA recovery is significantly less efficient.
HMW gDNA Purification Consists of Two Stages:
PART 2: HMW gDNA Binding and Elution
PART 1: Cell Lysis
- Pellet cells in a Monarch 2 ml Tube by centrifugation for 3 minutes at 1,000 x g. Frozen pellets should be thawed.
- Prepare Nuclei Prep and Lysis Solutions as indicated below:
a. Nuclei Prep Solution: Combine cold Nuclei Prep Buffer and RNase A according to the table below and vortex to mix. Keep on ice.STANDARD INPUT
LOW INPUT
# of SAMPLES
VOLUME OF NUCLEI PREP BUFFER (µl)
VOLUME OF RNase A (µl)
VOLUME OF NUCLEI PREP BUFFER (µl)
VOLUME OF RNase A (µl)
1
165
5.5
55
2
2
330
11
110
4
3
495
16.5
165
6
4
660
22
220
8
5
825
27.5
275
10
b. Nuclei Lysis Solution: Combine Nuclei Lysis Buffer and Proteinase K according to the table below and vortex to mix. Keep at room temperature.STANDARD INPUT
LOW INPUT
# of SAMPLES
VOLUME OF NUCLEI LYSIS BUFFER (µl)
VOLUME OF PROTEINASE K (µl)
VOLUME OF NUCLEI LYSIS BUFFER (µl)
VOLUME OF PROTEINASE K (µl)
1
165
11
55
4
2
330
22
110
8
3
495
33
165
12
4
660
44
220
16
5
825
55
275
20
- Flick to resuspend cell pellet. Add 150 µl (Low Input: 50 µl) of Nuclei Prep Solution and pipette up and down 10 times to mix, being careful not to introduce air bubbles. Incubate at room temperature for 2 minutes. The sample will become less turbid, indicating cell lysis; nuclei remain intact.
- Add 150 µl (Low Input: 50 µl) of Nuclei Lysis Solution to sample and invert 10 times to mix. Avoid introducing air bubbles. Do not vortex or pipette.
- Incubate at 56°C for 10 minutes in a thermal mixer with agitation at the desired speed to control the shearing and tune the size of gDNA. The speed of the thermal mixer influences fragment length; higher speeds reduce overall size. For the standard ligation-based Oxford Nanopore Technologies (ONT) sequencing protocol, agitation at 2,000 rpm is recommended. At 300 rpm, or with no shaking, maximal fragment length, in the Mb range, will be obtained (UHMW DNA). These samples will be highly viscous and difficult to process. Optimization may be required depending on the quality of the starting sample. Refer to “Choosing Agitation Speed During Lysis” for guidance. If desired, samples can be stored at 4°C overnight after the incubation.
- If working with multiple samples, prepare and label the plastics for Part 2: HMW gDNA Binding and Elution. Each sample will require:
- 1 Monarch Collection Tube II (no need to label).
- 1 Monarch Bead Retainer inserted into the collection tube; this will be used to remove the wash buffer from the gDNA bound to the beads.
- 1 Monarch 2 ml Tube; this will be used for eluting the gDNA from the beads.
- 1 1.5 ml microfuge tube (DNA low bind is recommended, not provided); this will be used to collect the eluate.
- Add 75 µl (Low Input: 25 µl) of Precipitation Enhancer after the 10-minute incubation and mix by inverting 8–10 times.
PART 2: HMW gDNA Binding and Elution
- Using clean forceps, add 2 DNA Capture Beads to each sample, which should be contained in a Monarch 2 ml Tube.
- Add 275 µl (Low Input: 100 µl) isopropanol, close the cap, and mix on a vertical rotating mixer at 10 rpm for 4 minutes to attach DNA to the beads. When working with ≥ 5 x 106 cells, double the inversion time to 8 minutes; this is especially important if low agitation speeds were used during lysis.
If a vertical rotating mixer is not available, invert slowly and gently by hand 30 times. A manual inversion is complete when the tube returns to the upright position. Slow inversion is critical for the DNA to bind to the beads; each full inversion should take ~5–6 seconds. If necessary, flick the tube to release any beads that stick to the bottom of the tube.
After a few inversions, the solution becomes more viscous and the DNA will wrap loosely around the beads. During the following inversions, precipitation of gDNA may be visible, especially with sample inputs ≥ 5 x 106 cells. The DNA complex will often contain small air bubbles. With more inversions, the DNA will completely wrap around the beads, often causing the beads to stick together.
- Remove and discard liquid by pipetting. Avoid removing any of the gDNA wrapped around the glass beads. For optimal DNA solubility, avoid letting the bound DNA dry out on the beads during this and the following steps; add the next buffer quickly. There are two suggested options for carrying out this step:
- Keeping tube upright, insert pipette tip and gently push beads aside to remove liquid.
- Angle the tube so that beads remain at the bottom, and liquid reaches toward tube opening. Pipette from the liquid surface and continue to angle as liquid is removed (tube will be almost horizontal at the end).
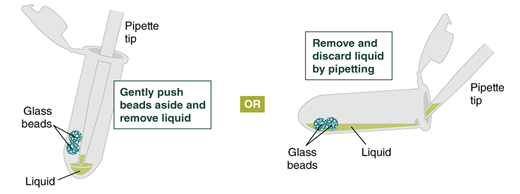
- Keeping tube upright, insert pipette tip and gently push beads aside to remove liquid.
- Add 500 µl gDNA Wash Buffer, close the cap, and mix by inverting the tube 2–3 times. Remove the gDNA Wash Buffer as described in step 3. The loose gDNA complex will condense around the beads more tightly.
- Repeat the wash in Step 4. Remove the gDNA Wash Buffer by pipetting. Alternatively, the buffer can be removed by decanting: position a pipette tip at the top of the angled tube to prevent the beads from falling out. It is not necessary to remove all the gDNA Wash Buffer at this point.
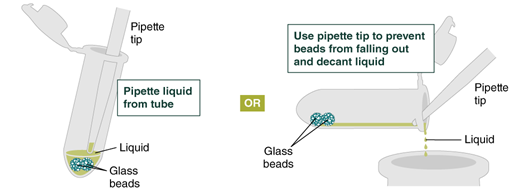
- Place a labeled bead retainer into a Monarch Collection Tube II. Pour the beads into the bead retainer and close the cap. Discard the used Monarch 2 ml Tube. When working with multiple samples, be sure to close the cap of the bead retainer after each transfer of beads.
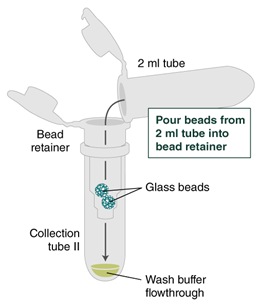
- Pulse spin (1 second or less) the sample in a benchtop minicentrifuge to remove any residual wash buffer from the beads.
- Separate the bead retainer from the collection tube, pour the beads into a new, labeled Monarch 2 ml Tube, and insert the used bead retainer into the labeled 1.5 ml microfuge tube (DNA low bind recommended, not provided) for later use during elution. Discard the used collection tube.
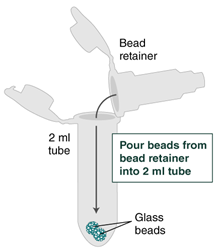
- Immediately add 100 µl (use 200 μl if working with > 2 x 106 cells) Elution Buffer II onto the glass beads and incubate for 5 minutes at 56°C in a thermal mixer with agitation at the lowest speed (300 rpm). Halfway through the incubation, ensure the beads are not stuck to the bottom of the tube by tilting the tube almost horizontally and gently shaking. This ensures that the beads can move freely, allowing for optimal release of the DNA from the beads. It also ensures that the lower bead does not stick to the bottom of the tube during the following transfer step. Elution volume can be reduced to as low as 50 μl without affecting recovery. However, if using < 100 μl, the gentle shaking of the sample should be done several times during the incubation to ensure complete wetting of the beads.
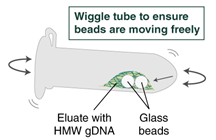
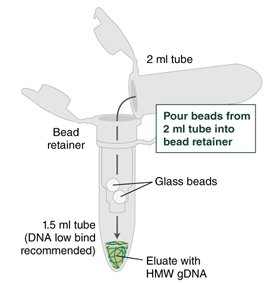
- Ensure the bead retainer is inserted into the 1.5 ml microfuge tube. Pour the eluate and the glass beads into the bead retainer and close the cap. When working with more than 1 sample, it is important to close the cap after each transfer of beads. Typically, all the eluate flows into the bead retainer upon pouring. If any volume remains in the 2 ml tube, spin briefly and transfer.
- Centrifuge for 30 seconds (1 minute if working with ≥ 5 x 106 cells) at 12,000 x g to separate the eluate from the glass beads. Discard the beads and retainer.
- Pipette eluate up and down 5–10 times with a wide bore pipette tip and ensure any visible DNA aggregates are dispersed. Before analysis or downstream use, HMW DNA must be homogeneously dissolved. After pipetting, incubate at 37°C for 30-60 minutes, overnight at room temperature, or for > 24 hours at 4°C. Pipette up and down 5-10 times again before analyzing or using the HMW DNA. Samples processed using low agitation speeds during lysis will require additional time to fully dissolve. See additional guidance in “Homogenization of HMW DNA Samples”. Samples can be stored at 4°C for short term use (weeks), or at -20°C for long term storage. The elution buffer (10 mM Tris, pH 9.0, 0.5 mM EDTA) is formulated for long term storage of gDNA.