EnGen® sgRNA Synthesis Kit, S. pyogenes Protocol (NEB #E3322)

Target-specific oligo design
This kit contains the S. pyogenes Cas9 Scaffold Oligo within the EnGen 2X sgRNA Reaction Mix, S. pyogenes. Target-specific oligos are designed by the user as follows:
- Select 20 nucleotide target sequence (not including the PAM (NGG) sequence). Use of a target DNA selection program is recommended. We recommend ChopChop.
- Check input sequence for presence of "G" at the 5´ end. If there are no "G's" at the 5´ end, add one "G" (making it a total of at least one G at the 5´ end).
- To the 5´ end; append T7 promoter sequence: TTCTAATACGACTCACTATA
- To the 3´ end; append 14 nucleotide overlap sequence: GTTTTAGAGCTAGA
- Check complete oligo sequence: 5´ TTCTAATACGACTCACTATAG(N)20GTTTTAGAGCTAGA 3´

sgRNA synthesis protocol
We strongly recommend wearing gloves and using nuclease-free tubes and reagents. Reactions should be assembled in microfuge tubes or PCR strip tubes.
- Thaw EnGen 2X sgRNA Reaction Mix, S. pyogenes, 0.1 M DTT and customer-supplied target-specific oligo (1 µM). Mix and pulse-spin each component in a microfuge prior to use. Store enzyme mix on ice but assemble reaction at room temperature.
-
Assemble the reaction at room temperature in the order listed. Avoid master mixes, and add the enzyme last to each reaction:
REAGENT AMOUNT Nuclease-free water
2 μl
EnGen 2X sgRNA Reaction Mix, S. pyogenes
10 μl
Target-specific DNA Oligo (1 µM)
5 μl
DTT (0.1 M) 1 μl EnGen sgRNA Enzyme Mix
2 μl
Total volume
20 μl
- Mix thoroughly and pulse-spin in a microfuge. Incubate at 37°C for 30 minutes.
- Transfer reaction to ice.
- For DNase treatment bring volume to 50 μl by adding 30 μl of nuclease-free water. Add 2 μl of DNase I (RNase-free, provided), mix and incubate at 37°C for 15 minutes.
- Proceed with purification of RNA or analysis by gel electrophoresis.
NOTE: sgRNAs synthesized using in vitro transcription methods from a DNA template and a bacteriophage polymerase will possess a 5´ triphosphate.
Optional: To remove the 5´ triphosphate (leaving a 5´ OH), treatment of the sgRNA with Alkaline Phosphatase, Calf Intestinal (CIP) (NEB #M0290) can be performed. Phenol:chloroform extraction and/or cleanup by spin column is recommended to inactivate and remove the phosphatase.
Purification of sgRNAs
Spin columns will remove proteins, salts and most unincorporated nucleotides. Please ensure that spin columns are compatible with the size of sgRNAs (~100 nts) and expected RNA yields (4–25 µg). Follow manufacturer’s instructions. We recommend Monarch RNA Cleanup Kit (50 μg) (NEB #T2040) for purification .
Evaluation of sgRNA transcription products
Quantitation by UV Light Absorbance
sgRNA concentration following purification can be determined by measuring absorbance at 260 nm on a standard UV spectrophotometer following dilution of the sample, or read directly on a NanoDrop® (recommended). 1 OD260 unit for single-stranded RNA (ssRNA) is 40 µg/ml. RNA concentration can be determined as follows:
A260 x dilution factor x 40 = "x" µg/ml ssRNA.
Expected yields following purification can be 4–25 µg.
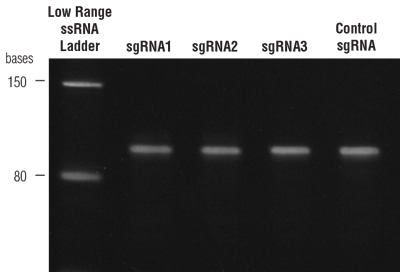
Analysis of sgRNAs by gel electrophoresis
- Assessment of sgRNA quality and length can be evaluated by gel electrophoresis. Gels should be run under denaturing conditions to avoid formation of secondary structure.
- Mix an aliquot of RNA with RNA Loading Dye, (2X) (NEB #B0363) or Urea Dye.
- Denature RNA according to manufacturer’s recommendation and load onto a TBE-urea gel, such as Novex® TBE-urea gels.
- Visualize RNA by staining the gel with SYBR® Gold or ethidium bromide.
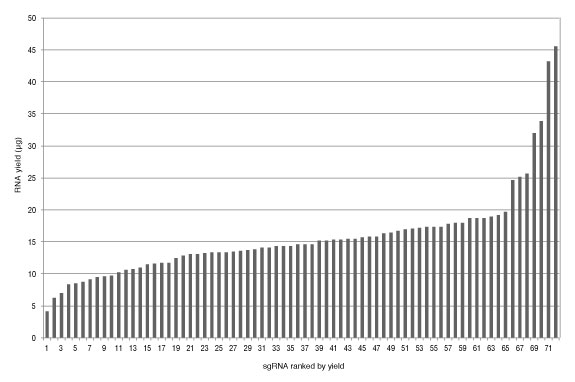
In-vitro Cas9 Nuclease Assay
S. pyogenes Cas9 double-stranded DNA nuclease is guided to the target DNA based on sequence complementarity to the sgRNA that is loaded into the protein. Site-specific DNA cleavage occurs three nucleotides upstream of the PAM sequence (NGG), within the complementary region. sgRNAs synthesized with this kit can be used with Cas9 nuclease, S. pyogenes (NEB #M0386) or EnGen Spy Cas9,NLS (NEB #M0646) for in vitro cleavage reactions.