Glycomics:
A rapidly evolving field with a sweet future
by Christopher H. Taron1 and Pauline M. Rudd2
1 – New England Biolabs, Ipswich MA 2 – NIBRT GlycoScience Group, National Institute for Bioprocessing Research and Training, Dublin, Ireland and BTI, A*STAR, Singapore
Glycobiology is entered in the Oxford English Dictionary as “f. GLYCO- + BIOLOGY n.: coined by Prof. Raymond Dwek in 1982” and is defined as the branch of science concerned with the role of sugars in biological processes. Glycobiology addresses the assembly, structure and biology of chains of sugars (termed ‘glycans’). Glycans are widely distributed in nature and have physical, chemical, and biological properties that make them important players in areas such as biofuels, food, materials science, biotechnology, and pharmaceuticals. Glycans are also among the most important molecules in cell biology. Together with nucleic acids, proteins, and lipids, glycans are one of the four basic building blocks from which all cells are comprised (1).
Introduction
Glycans play many critical roles in both the normal function of cells and in disease. They assist in the folding of many proteins, aid in protein trafficking, mediate cell adhesion, differentiate blood groups, modulate the immune system, are implicated in many signaling pathways, and provide a protective extracellular matrix for many types of cells. Glycans are also implicated in the process of infectivity for many pathogenic bacteria (2) and most viruses (3), including those that cause the common cold, influenza, and HIV/AIDS.
Individual glycans are assembled from monosaccharides that are linked together via glycosidic bonds, and can be covalently bound to various proteins and lipids (termed ‘glycoconjugates’ in this context). Several classes of glycoconjugates are synthesized by mammalian cells (Figure 1) and populate the membranes of the secretory pathway, the cell surface, and the extracellular matrix. Glycans that are appended to certain serine/threonine residues (O-glycans) or certain asparagine residues (N-glycans) of secretory proteins are the most abundant post-translational modifications of proteins. It is estimated that >50% of mammalian proteins possess appended glycans (4) and the surface of each mammalian cell may contain as many as 10 million N- or O-linked glycans attached to proteins (5). In addition, N- and O-linked glycans are present on nearly all proteins that are secreted from cells. Thus, glycoproteins are present in all mammalian body fluids.
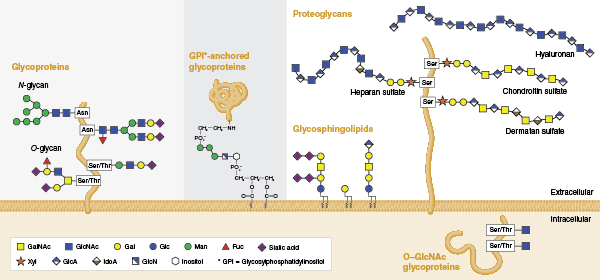
Glycan structure is inherently complex, a fact that relates to the way glycans are synthesized. Unlike DNA or proteins that are built from copying or interpreting a genetic template, glycans are assembled through the action of complex biosynthetic pathways. The flexibility built into this system allows cells to alter their glycan structures to respond rapidly to changes in their environment without needing to alter their genomes. It is estimated that the human genome encodes over 900 proteins involved in various aspects of glycan assembly or recognition (6). Of these, more than 200 human genes encode glycosyltransferases, enzymes that specifically add new sugars to glycans. The expression of many of these proteins can vary amongst different cell types and tissues giving rise to significant glycan structural variation. For example, over 140 structural variants of N-glycans alone have been identified on glycoproteins present in human serum (7). In addition, for any individual mammalian glycoprotein, numerous different glycans can be attached and processed. A glycoform is a single copy of a glycoprotein with a defined glycan present at each glycosylation site. Thus, proteins exist as collections of glycoforms and the sugars that are present depend on the cell, the protein, and the 3D structure around the site of their attachment.
In this article, we give an overview of the emergence of high-throughput glycomics and its potential to improve our understanding of the roles glycans play in cell biology. We will also summarize the rapidly growing field of mammalian serum glycomics and discuss its potential as a new reservoir of biomarkers of health and disease, and its diagnostic potential.
Technical Advances in Glycomics
Over the past two decades, the cellular and molecular biology fields have been transformed by ‘omics’ technologies that permit analysis of large pools of biological molecules en masse. The advent of deep sequencing technologies has dramatically changed the analysis of nucleic acids and has revolutionized the fields of genomics and transcriptomics. Additionally, improvements in mass spectrometry sensitivity and speed has strongly impacted the depth to which proteomes can be explored. However, nucleic acids and proteins do not alone provide a complete view of cellular function. Enabled by similar recent analytical advances, the past decade has also seen rapid growth of glycomics, lipidomics and metabolomics, fields that explore cellular compounds that are not synthesized by genetic template-driven processes, but instead are products of biosynthetic pathways created by a cell’s proteome. These newer fields, in conjunction with the more mature fields of genomics, proteomics and transcriptomics, are now generating a more holistic view of cellular function during both normal cell physiology and in disease (Figure 2).
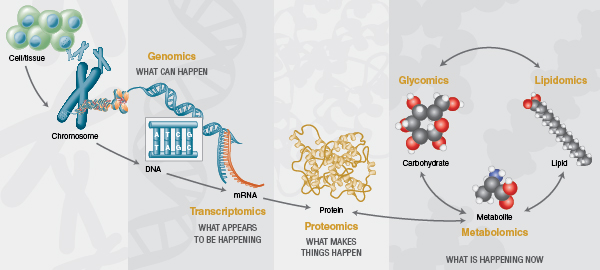
Glycomics addresses the comprehensive high-throughput structural analysis of glycans produced by an organism or its components (e.g., cells, organs, body fluids, etc). The number of published glycomics studies has been steadily increasing over the past decade. This growth has been fueled by technical advancement in nearly every aspect of glycan analysis (see ref. 8 for a recent technical review). New enzymatic tools have emerged that permit faster and/or more complete liberation of glycans from proteins (9) and glycolipids (10). Highly pure recombinant versions of the field’s preferred exoglycosidases are now available, and the first array consisting of pre-mixed combinations of these enzymes optimized for high-throughput glycan sequencing applications has been devised by New England Biolabs. New fluorescent dyes have been developed with advantages for more quickly labeling glycans and/or increased sensitivity in mass spectrometry applications (9). Improvements have been made to solid-phase extraction techniques and faster chromatographic separation of glycan samples using ultra-high performance liquid chromatography systems or capillary electrophoresis. Bioinformatics resources such as GlycoBase (11, 12), a database that correlates known glycan structures to their liquid chromatography mobilities, have been developed to improve structural interpretation of glycan analysis data. Finally, high-throughput analyticalworkflows that incorporate many of these innovations have been devised to expand the applicability of glycomics to large sample sets (10, 13, 14). Armed with these new analytical tools, researchers have begun unlocking the exciting potential of the mammalian glycome.
Mammalian Serum Glycomics
One area of the glycomics field that has seen remarkable progress over the past decade is the characterization of the mammalian blood serum glycome using N-glycan profiling. In this field, N-glycans are isolated en masse from glycoproteins that circulate in the mammalian bloodstream (or commonly from IgG purified from serum), and are subjected to chromatography and structural profiling (Figure 3). Serum N-glycan profiles from different cohorts of individuals (e.g., healthy versus disease samples) can be compared to identify changes in the abundance of individual glycan species.
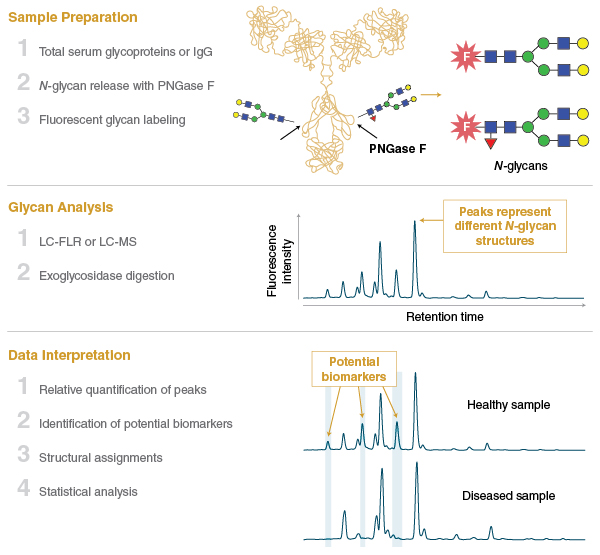
This methodology has been transforming our understanding of the serum glycome and how it changes in response to both normal physiological processes like aging, or the onset of various diseases such as cancer. Features of the serum and IgG glycomes have already been shown to be associated with genome, transcriptome, and metabolome changes in individuals, and point the way to a more detailed understanding of disease pathways (15–19). In the coming years, there is amazing potential for the serum glycome to be exploited as a predictor of health and disease, and to have a role in personalized medicine.
Serum N-glycan profiling has already identified glycan biomarkers associated with the aging process. Studies have shown that our glycomes change in predictable ways as we grow older or when our bodies undergo major physiological transitions, such as adolescence or menopause in women (20). Serum glycan biomarkers have also been used to estimate both chronological age (one’s actual age) and biological age (a measure of how well a body is performing relative to its chronological age) (21). Because we all age at different rates due to differences in physiology and lifestyle choices, it is possible that serum glycans may someday be used to help manage each individual’s personal health risks associated with the onset of age-related illnesses. Finally, glycan biomarkers have been used to estimate human age from N-glycan profiles obtained from dried bloodstains (22), suggesting that analysis of blood N-glycans may also find application in forensic medicine. The serum N-glycome can also predictably change in response to disease. Correlations between changes in the composition and/or abundance of circulating N-glycans and many diseases have been observed. A noN-exhaustive list includes numerous cancers (e.g., breast, gastric, liver, lung, pancreatic, ovarian and prostate), rheumatoid arthritis, and schizophrenia (23–25). Additionally, compositional analysis of glycans has been used to identify glycan markers that correlate to good and poor prognoses in cohorts of prostate cancer patients (25) and breast cancer patients (19), suggesting that glycan profiling could someday have potential in patient stratification and disease staging. Altered N-glycan profiles have also been observed in serum from mammals affected by infectious diseases such endometritis (a postpartum bacterial infection in cattle; 26), Leishmania parasite infection in humans (27), and Dirofilaria immitis (heart worm) infection in dogs (28).
Serum Glycans as Diagnostics
There is hope that serum glycans will help change the way in which many diseases are diagnosed and/or monitored during treatment. In the field of genomics, there has been much discussion around the promising concept of “liquid biopsy” (a blood test) for the diagnosis of cancer without the need for invasive tissue biopsy (see ref. 29 for a recent review). In this approach, tumor cells or tumor cell DNA circulating in the bloodstream of patients with certain cancers can be isolated and subjected to genomic sequencing to identify mutations that give information as to how a tumor is changing. Serum glycomics has the potential to further expand upon the promise of the liquid biopsy concept. Both analyses seek to detect tumors earlier, characterize the type or stage of a cancer, or determine how a cancer is responding to treatment, directly from small amounts of patient’s blood instead of invasive tumor biopsy.
One hope is that serum glycan biomarkers might live up to the published data and be more valuable as earlier indicators of disease than current methods. This stems from several observations about the blood glycome. First, in contrast to tumor-derived nucleic acids, glycans are very abundant molecules in serum. For example, glycan profiles can be obtained from as little as 5 microliters of serum. Secondly, changes in serum glycan structures may be more immediate indicators of a changing physiology. For example, some changes in the serum glycome composition could arise from the increasing presence of many aberrantly glycosylated proteins secreted by tumor cells. Alternatively, changing glycosylation patterns on circulating immune system proteins may indicate a systemic response to a tumor or infectious agent’s presence (i.e., the types of changes currently being observed using IgG N-glycan profiling). In the latter case, the serum IgG glycome may act as an early sensor that a problem is developing or changing.
Some glycan biomarker studies support the notion of earlier detection of disease and physiological changes. For example, diagnostic glycan biomarkers of bovine endometritis are detectable in the maternal bloodstream as early as seven days after birth of a calf (prior to the onset of clinical symptoms) compared to diagnosis upon presentation of physical signs of infection at 2-5 weeks postpartum (26). Similarly, bovine pregnancy can be predicted from a glycan biomarker present in a cow’s milk some 2-4 weeks earlier than the standard method of detection of pregnancy by rectal ultrasound (26). The field of serum glycan biomarker discovery has made tremendous progress, however, it remains to be determined if it will inspire a new generation of clinical diagnostics. It is conceivable that serum glycan profiling could provide an orthogonal blood-based analysis to DNA sequencing, or other diagnostic methods, to increase the statistical confidence around early tumor identification orprogression. This could help inform the best choices for treatment. Such “liquid biopsy” concepts will require significantly more research in the coming years to determine their clinical feasibility.
Perspective and Conclusions
The past decade has seen tremendous progress in the field of glycomics. Enabled by numerous technical advances in analytical sensitivity and throughput, researchers are now starting to unlock the secrets of the mammalian glycome. Glycomics is now being used in combination with genomics, epigenomics, proteomics, lipidomics and metabolomics to provide a more holistic view of how cellular pathways function and how they change in response to disease. In the field of human serum glycomics, there has been an explosion in strong associations made between changes in glycan structure and various diseases, disease stages and traits, changes in circulating tumor cells, and age-related changes to our physiology. These correlations are revealing critical new insights into how diseases develop.
While much progress has been made, there is still a mountain to climb. The coming years will continue to see improvements to glycan analytical techniques with emphasis on analytical automation and miniaturization. Additionally, bioinformatics will play an increasing role in improving the interpretation of glycomics data. To date, the lessons learned about the human serum glycome have come largely from the analysis of N-glycans. The field will strive to expand serum glycomics into other classes of circulating glycans such as glycosphingolipid head group glycans (10) and O-glycans. Finally, it is anticipated that the first glycaN-based diagnostic tests will start to emerge. It is clear that it is an exciting time for glycomics. The field sits in a position to write an important chapter in our understanding of cell biology and positively affect the direction of human health.
Acknowledgments
The authors thank Prof. Stephen Carrington and Prof. Mark Crowe of University College Dublin, Ireland for sharing unpublished information on glycan biomarkers in cattle. The authors also thank Andrew Bertera, Alicia Bielik, Rebecca Duke, Mehul Ganatra, Ellen Guthrie, Jeremy Foster, Paula Magnelli, Beth McLeod, Stephen Shi, Barbara Taron, and Saulius Vainauskas for comments on the article.
References
- Marth, J.D. (2008) Nat. Cell Biol. 10, 1015-1016.
- Day, C.J., et al. (2015) Proc. Natl. Acad. Sci. USA, 112, E7266–E7275.
- Ströh, L.H. & Stehle, T., (2014) Annu. Rev. Virol. 1, 285-306.
- Apweiler, R., et al. (1999) Biochim. Biophys. Acta. 1473, 4-8.
- Wang, Z., et al. (2013) Science, 341, 379-383.
- Cummings, R.D. & Pierce, J.M. (2014) Chem. Biol. 21, 1-15.
- Saldova, R., et al. (2014) J. Proteome Res. 13, 2314-2327.
- Duke, R.M. & Taron, C.H. (2015) Biopharm International, 28, 59-64.
- Lauber, M.A., et al. (2015) Anal. Chem. 87, 5401-5409.
- Albrecht, S., et al. (2016) Anal. Chem. 88, 4795-4802.
- Campbell, M.P., et al. (2008) Bioinformatics, 24, 1214-1216.
- Campbell, M.P., et al. (2015) Methods Mol. Biol. 1273, 17-28.
- Stöckmann, H., et al. (2013) Anal. Chem. 85, 8841-8849.
- Stöckmann, H., et al. (2015) Integr. Biol. (Camb), 7, 1026-1032.
- Butler, M., et al. (2003) Glycobiology, 13, 601-622.
- Lauc, G, (2010) PLoS Genet. 6, e1001256.
- Igl W., et al. (2011) Mol Biosyst. 7, 1852-1862.
- Lauc, G., et al. (2013) PLoS Genet. 9, e1003225.
- Haakensen, V.D., et al. (2016) Mol Oncol. 10, 59-72.
- Knezevic, A. et al. (2010) Glycobiology, 20, 959-969.
- Kristic, J., et al. (2014) J. Gerontol. A. Biol. Sci. Med. Sci. 69, 779-789.
- Gudelj, I., et al. (2015) Int. J. Legal Med. 129, 955-961.
- Telford, J.E ., et al. (2011) Biochem. Soc. Trans. 39, 327-330.
- Adamczyk, B., et al. (2012) Biochim. Biophys. Acta. 1820, 1347-1353.
- Ruhaak, L.R., et al. (2013) Mol. Cell. Proteomics, 12, 846-855.
- Carrington S. & Crowe, M., University College Dublin, personal communication.
- Gardinassi, L.G., et al. (2014) mBio 5, e01844-14.
- Duke, R.M., Petralia, L., Taron, C.H. & Foster, J.M., unpublished observations.
- Karachaliou, N., et al. (2015) Ann. Transl. Med. 3, 36.
Additional NEB Resources:
- Redesigning glycosidase manufacturing quality for pharmaceutical and clinical applications
- Products available from NEB for Glycomics research
- NEB’s PNGase F – Designed for your Application

