Choice of One-Step RT-qPCR or Two-Step RT-qPCR
Return to DNA Amplification, PCR and qPCRRT-qPCR (Reverse Transcription Quantitative PCR) is a valuable tool for assessing gene expression by measuring the abundance of certain RNAs. Two general methods are available: one-step RT-qPCR and two-step RT-qPCR. In both cases, RNA is first reverse-transcribed into cDNA, which is then used as the template for qPCR amplification. In one-step RT-qPCR, cDNA synthesis and qPCR are performed in a single reaction vessel in a common reaction buffer. In two-step RT-qPCR, cDNA is synthesized in one reaction, and an aliquot of the cDNA is then used for a subsequent qPCR experiment.
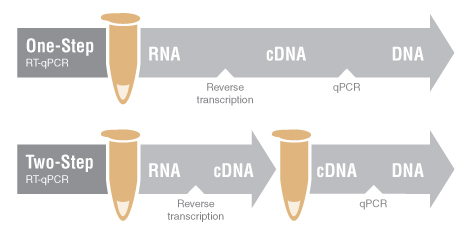
One-Step RT-qPCR
In one-step RT-qPCR, cDNA synthesis and qPCR are carried out in the same reaction vessel, in a common buffer. Gene-specific primers direct cDNA synthesis and amplification of a specific target. Because specific primers typically anneal at higher temperatures than random primers, one-step protocols often use higher RT reaction temperatures than two-step workflows and employ engineered or novel RTs that can tolerate higher reaction temperatures. Major advantages of one-step reactions include minimal sample handling, reduced bench time, and closed-tube reactions, reducing chances for pipetting errors and cross-contamination. This makes one-step an especially strong choice for quantitating the same gene(s) repeatedly, particularly in high-throughput applications and in diagnostic settings.
The quality and scarcity of RNA samples are important considerations for one-step RT-qPCR. Where RNA is of low quality, and inhibitors are inconsistently present between samples, reaction efficiency can be impacted to varying degrees, reducing comparability between samples. In addition, because the cDNA synthesis product cannot be saved after one-step RT-qPCR, additional aliquots of the original RNA sample(s) are required in order to repeat reactions or to assess the expression of other genes.
Two-Step RT-qPCR
In two-step RT-qPCR, cDNA synthesis is carried out using random hexamers, oligo-dT primers, and/or gene-specific primers. Reactions using random hexamers and/or oligo-dT primers produce cDNA that is a mixture of various RNA species in the sample. The cDNA synthesis reaction can be scaled up to accommodate higher RNA input, and extraction and precipitation steps can be used to concentrate and/or further purify the cDNA. This allows better control over the amount of cDNA template that is used for qPCR quantitation. Because only a portion of the cDNA product is used in the subsequent qPCR step, remaining cDNA can be stored for future use, or quantitating the expression of multiple genes from a single RNA/cDNA sample.
Two-step RT-qPCR uncouples cDNA synthesis and subsequent qPCR, allowing greater freedom in selecting RT enzymes and qPCR reagents separately. This flexibility can be useful for controlling sequence representation, efficiency, and optimization of difficult RT-qPCR reactions.
The drawback of two-step RT-qPCR is that it requires an extra open-tube step, more pipetting manipulations, and longer hands-on time. This can lead to greater variability and risk of contamination, and makes two-step RT-qPCR less amenable to high-throughput applications.
Table 1. Comparison of one‑step and two‑step RT‑qPCR
| RT Primers | Advantages | Disadvantages | Ideal uses | |
|---|---|---|---|---|
One-Step RT-qPCR |
|
|
|
|
| Two-Step RT-qPCR |
|
|
|
|
One-Step and two-Step RT-qPCR reagents available from NEB
One-Step RT-qPCR featuring Luna® WarmStart® Reverse Transcriptase- Dye-based detection: Luna® Universal One-Step RT-qPCR Kit (NEB #E3005)
- Probe-based detection: Luna Universal Probe One-Step RT-qPCR Kit (NEB #E3006)
- Dye-based detection directly from cells: Luna Cell Ready One-Step RT-qPCR Kit (NEB #E3030)
- Probe-based detection directly from cells: Luna Cell Ready Probe One-Step RT-qPCR Kit (NEB #E3031)
- Cell lysis module: Luna Cell Ready Lysis Module (NEB #E3032)
Two-Step RT-qPCR
- cDNA synthesis: For two-step applications, we recommend LunaScript® RT SuperMix Kit (NEB #E3010). NEB offers a variety of additional cDNA synthesis solutions. For more information, please consult the NEB cDNA Synthesis Selection Chart.
- Dye-based detection: Luna® Universal qPCR Master Mix (NEB #M3003)
- Probe-based detection: Luna® Universal Probe qPCR Master Mix (NEB #M3004)

