DNA Enrichment
Choose Type:
- Capture and Elute Step for Control DNA (E2600)
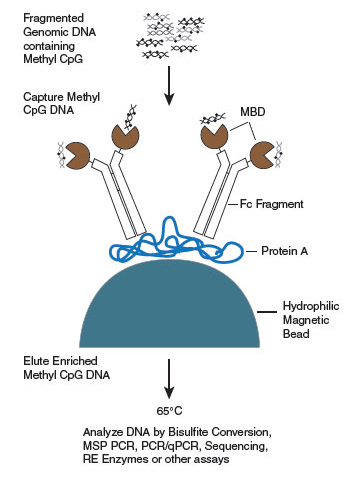
- Capture Methylated CpG DNA (E2600)
- Cross-linking of IgG to Protein A or G Beads
- DNA Fragmentation (E2600)
- Downstream Analysis (NEB #E2600)
- Elute Captured Methylated CpG DNA (E2600)
- Immunoprecipitation using Protein A/G Magnetic Beads
- Phage Display: Solution-phase Panning with Affinity Bead Capture
- Prebind MBD2a-Fc to Protein A Magnetic Beads (NEB #E2600)
- Purification of IgG using Protein A/G Magnetic Beads
- Wash Off Unbound DNA (E2600)
- Protocol for use with NEBNext ChIP-seq Library Prep Reagent Set for Illumina (E6200)
- Epigenetics Brochure
Brochures
- Wee E., Ngo T., Trau M. (2015) A simple bridging flocculation assay for rapid, sensitive and stringent detection of gene specific DNA methylation Sci Rep; 5, 15028. PubMedID: 26458746, DOI: 10.1038/srep15028
Products and content are covered by one or more patents, trademarks and/or copyrights owned or controlled by New England Biolabs, Inc (NEB). The use of trademark symbols does not necessarily indicate that the name is trademarked in the country where it is being read; it indicates where the content was originally developed. The use of this product may require the buyer to obtain additional third-party intellectual property rights for certain applications. For more information, please email busdev@neb.com.
This product is intended for research purposes only. This product is not intended to be used for therapeutic or diagnostic purposes in humans or animals.


