Site Directed Mutagenesis
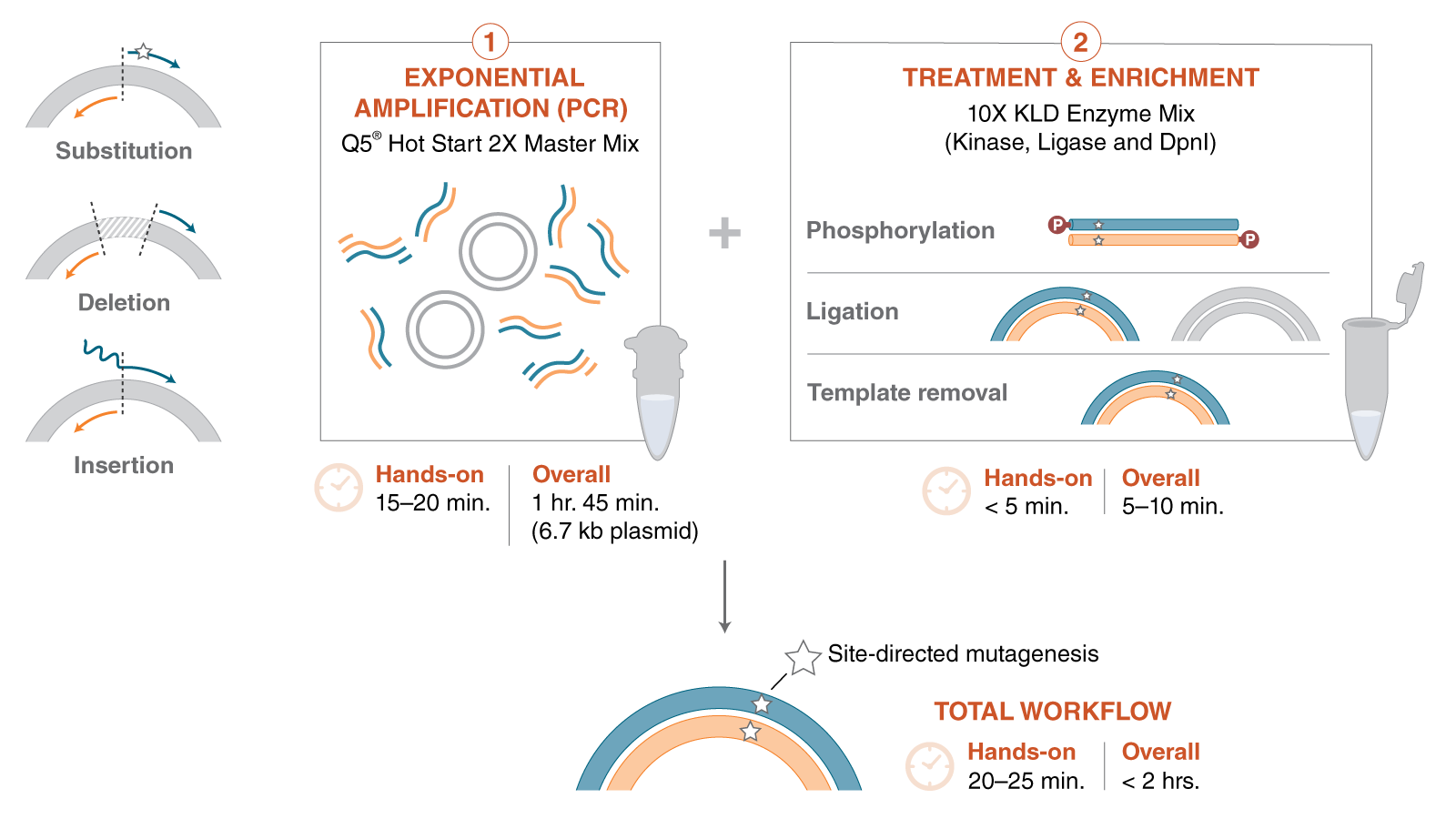
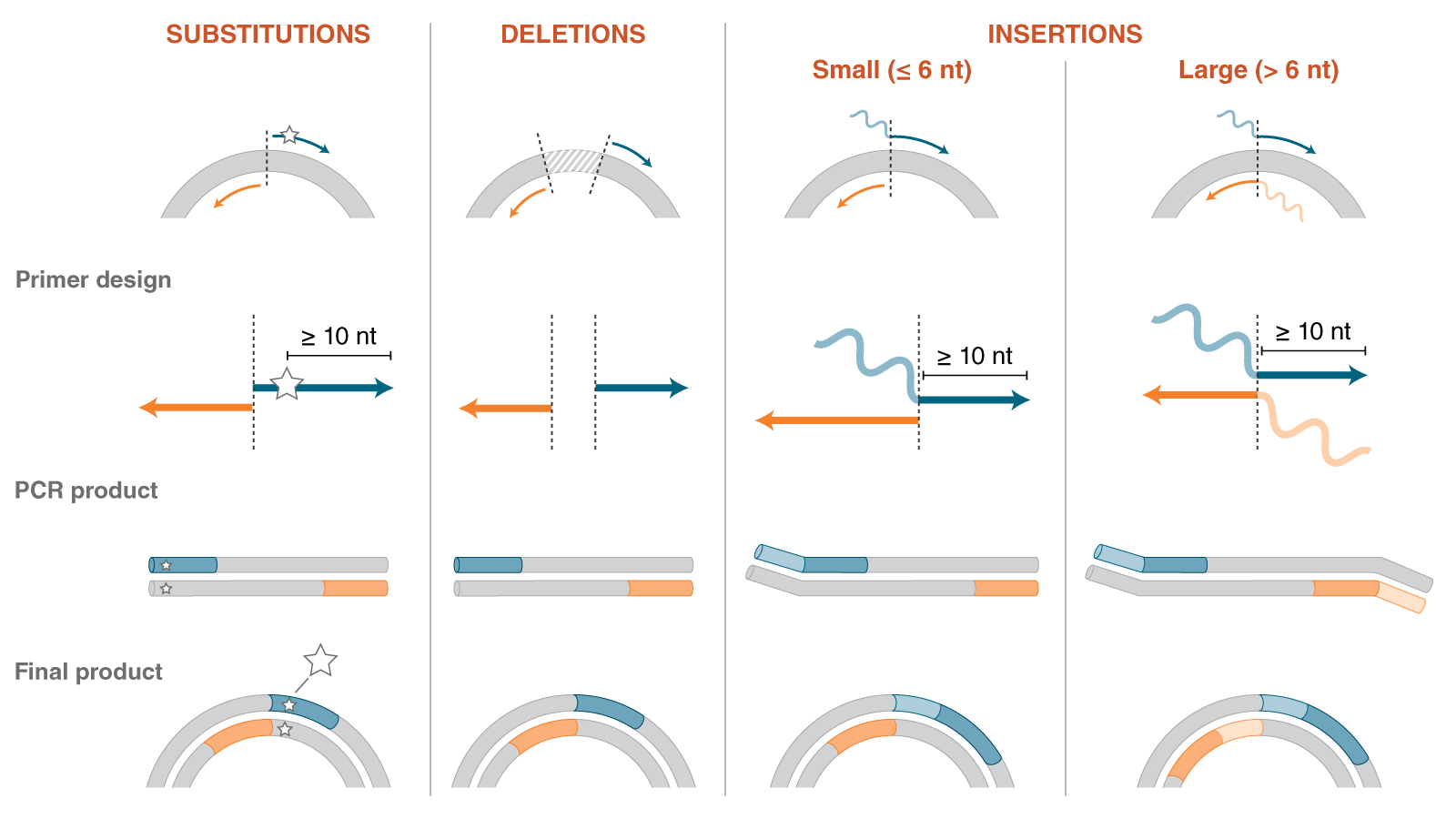
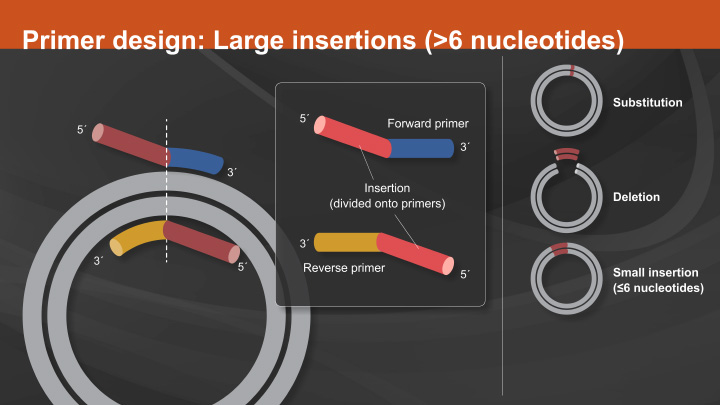
Site-directed mutagenesis (SDM) is a method to create specific, targeted changes in double stranded plasmid DNA. There are many reasons to make specific DNA alterations (insertions, deletions and substitutions), including:
- To study changes in protein activity that occur as a result of the DNA manipulation.
- To select or screen for mutations (at the DNA, RNA or protein level) that have a desired property
- To introduce or remove restriction endonuclease sites or tags
Choose Type:
- Protocol for Control Reaction (E0554)
- Protocol for Q5® Site-Directed Mutagenesis Kit (E0554)
- Quick Protocol for Q5® Site-Directed Mutagenesis Kit (E0554)
- Protocol for Control Reaction (E0552)
- Protocol for Q5® Site-Directed Mutagenesis Kit (Without Competent Cells) (E0552)
- Quick Protocol for Q5® Site-Directed Mutagenesis Kit (Without Competent Cells) (E0552)
- KLD Enzyme Mix Reaction Protocol (M0554)
- Molecular Cloning Technical Guide
- Troubleshooting Guide for Cloning
Brochures
Troubleshooting Guides
- Yafeng Li, Delu Song, Ying Song, Liangliang Zhao, Natalie Wolkow, John W Tobias, Wenchao Song, Joshua L Dunaief (2015) Iron-induced Local Complement Component 3 (C3) Up-regulation via Non-canonical Transforming Growth Factor (TGF)-β Signaling in the Retinal Pigment Epithelium. J Biol Chem; 290, 11918-34. PubMedID: 25802332, DOI: 10.1074/jbc.M115.645903
Products and content are covered by one or more patents, trademarks and/or copyrights owned or controlled by New England Biolabs, Inc (NEB). The use of trademark symbols does not necessarily indicate that the name is trademarked in the country where it is being read; it indicates where the content was originally developed. The use of this product may require the buyer to obtain additional third-party intellectual property rights for certain applications. For more information, please email busdev@neb.com.
This product is intended for research purposes only. This product is not intended to be used for therapeutic or diagnostic purposes in humans or animals.