Disulfide bond formation in the cytoplasm of SHUFFLE® cells (4 of 4)
Script
We will now look into disulfide bond formation in the cytoplasm.
The periplasm is an oxidative compartment, while the cytoplasm is a reducing compartment. This is due to two existing pathways: the thioredoxin pathway…and the glutaredoxin pathway. NADPH provides the major pool of electrons in the cytoplasm, supplying electrons to the two reducing pathways.
In the glutaredoxin pathway, glutathione reductase, or “gor”, reduces the enzymes GshA and GshB, which, in turn, synthesize glutathione. Glutathione then maintains the glutaredoxins in their reduced state, thereby reducing any disulfide bond formed.
In the thioredoxin pathway, thioredoxin reductase, or trxB, reduces thioredoxins, which, in turn, reduce any disulfide bond formed.
Glutaredoxins, together with thioredoxins, are the main reducing enzymes in the cytoplasm, and reduce any disulfide bond formed. These two pathways are redundant and are responsible for maintaining the proteome in a reduced state.
Knocking out one pathway does not lead to lethality, as the other pathway is sufficient for cell viability. Genetic knockouts of both gor AND trxB are lethal to the cells, as cells require a certain level of reducing capacity for viability.
Genetic selection of suppressors of trxB/gor lethality is mapped onto the peroxidase pathway. The peroxidase pathway receives its electrons from the pool of NADH, via peroxide reductase, AhpF. AhpF maintains the peroxidase AhpC in its active, reduced state. Wild type AhpC, which normally is responsible for the removal of peroxides, is mutated to AhpC*, which can now reduce glutaredoxins. This allows for sufficient reducing power for cell viability.
However, thioredoxins remain in their oxidized state, making them capable of forming disulfide bonds in the cytoplasm.
SHuffle cells lack trxB and gor, but the lethality is suppressed by AhpC*, resulting in disulfide bonds catalyzed by thioredoxins. However, unlike the periplasm, the cytoplasm lacks any disulfide bond isomerase essential for the folding of complex, multi-disulfide bonded proteins.
To increase the fidelity of disulfide bond formation, the periplasmic disulfide bond isomerase, DsbC, has been engineered to be overexpressed in the cytoplasm, allowing for the correct folding of multi-disulfide bonded proteins.
For certain multi-disulfide bonded proteins, SHuffle is the only E. coli strain that can express active protein.
This is illustrated when tissue plasminogen activator, with six disulfide bonds, is expressed in various strains. TPA is not active in wild-type cells, nor in cytoplasmically oxidative FA113 strains. However, TPA can fold and become active in the presence of cytoplasmic DsbC, when expressed in SHuffle cells.
The SHuffle cell lines are available in two versions: E. coli K12 and E. coli B, and are commercially available from NEB.
Related Videos
-
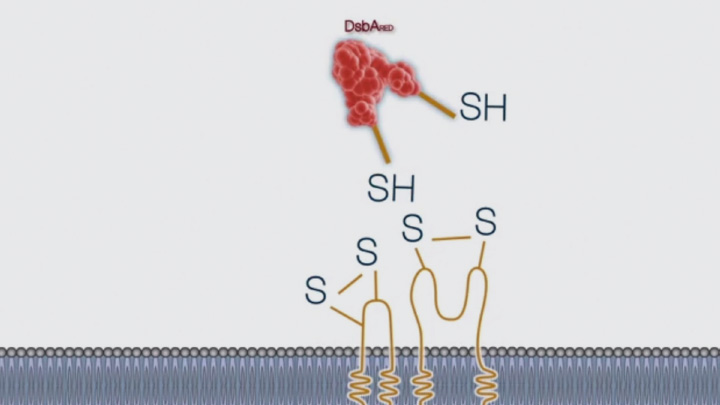
Disulfide bond formation in the periplasm of E. coli and the cytoplasm of SHUFFLE® -

Stages of SHUFFLE® Optimization -
What is a Disulfide Bond (1 of 4)

