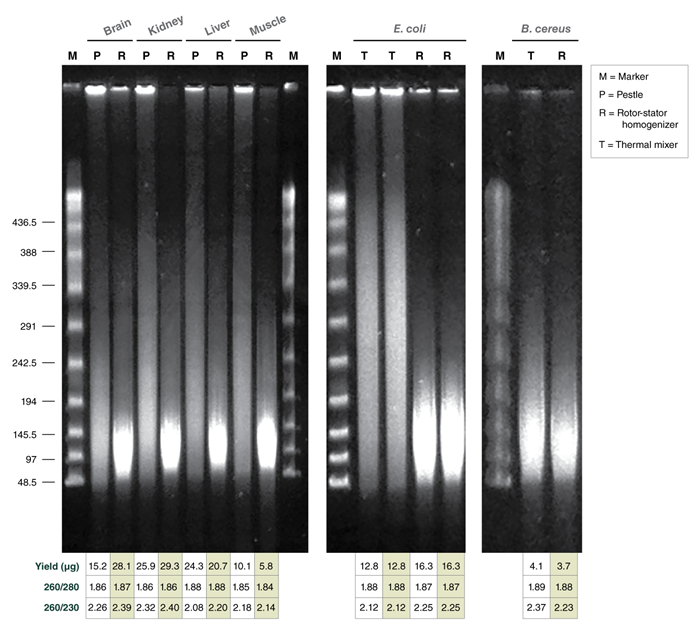
If the goal is to isolate ultra HMW DNA, microtube pestle homogenization is the best choice. Rotor-stator homogenizer (RSH) is an efficient alternative, but the HMW DNA that is released into the lysate may undergo a mild degree of shearing during the homogenization process. Therefore, for recovering the longest DNA fragments using the RSH, it should be used at the lowest setting and the device should be switched off immediately once the tissue pieces are completely disrupted. Generally, for pestle homogenization, the more thoroughly a sample is homogenized, the shorter the lysis time will be and the larger, more intact the DNA will be. The sample should be ground into the thinnest possible layer so that lysis by Proteinase K happens quickly. However, the tissue structure also influences the speed of lysis, and as a consequence, the fragment length. Soft organ tissue and brain will lyse in seconds and will give the longest fragments; muscle tissue may need a bit longer but should be lysed within a minute or so. Very rigid fibrous tissue (e.g., mouse tail) needs several minutes to lyse completely and therefore tends to give shorter DNA fragments. Since tissue cells cannot be freely resuspended as cultured cells can, portions of the cells lyse simultaneously, causing a degree of tangling of the HMW DNA. This tangling can be observed as upward smearing in a pulsed-field gel lane. Rotor-stator homogenizer treatment can cut those aggregates apart, for the most part. Pestle-treated samples however, particularly when frozen tissue is used, may require some needle shearing to resolve these DNA aggregates in order to get optimal long read sequencing results.