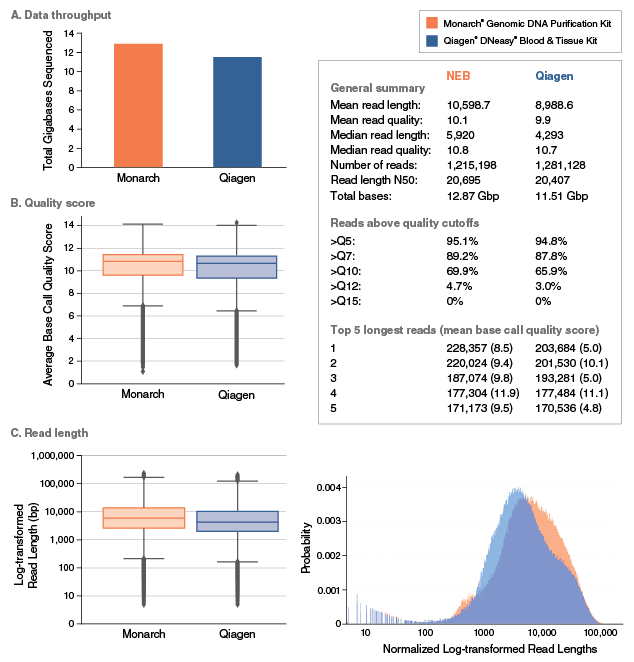
Yes. gDNA purified with this kit can be used for various applications on these sequencing platforms. Assuming high quality input material has been used, read lengths of up to 50 kb can be routinely achieved using Monarch purified gDNA from different starting materials.
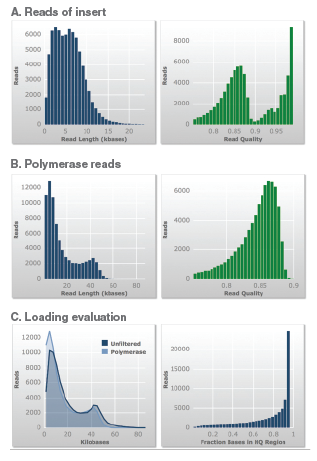
Total bases (yield): 1208 Mb, Average Polymerase read length: 16,046, Polymerase read quality: 0.85. Average Reads of Insert length: 6,060, Reads of Insert quality: 0.89, Longest read of insert: 43,680.
A. Insert lengths indicate that the majority of the reads were in the desired insert size range.
B. The polymerase read lengths indicate that the purified DNA was of high quality to allow the enzyme to read the insert several times within the SMRTbell construct.
C. The loading evaluation data further illustrates that the purified DNA is of high quality.

