Traditional Cloning Workflow
Script
Related Videos
-
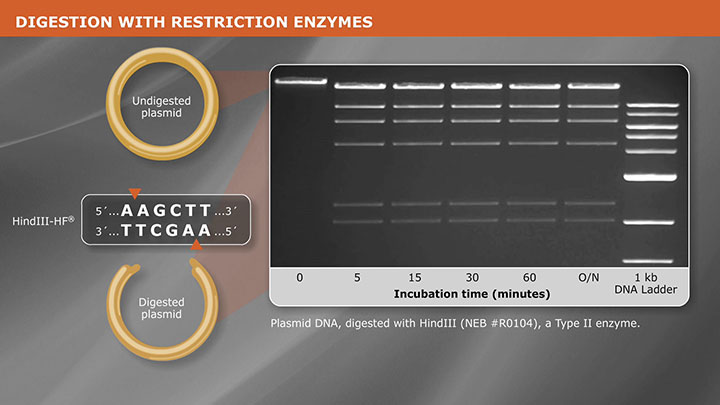
Cloning With Restriction Enzymes -
NEB® TV Ep. 8 – Cloning and DNA Assembly -
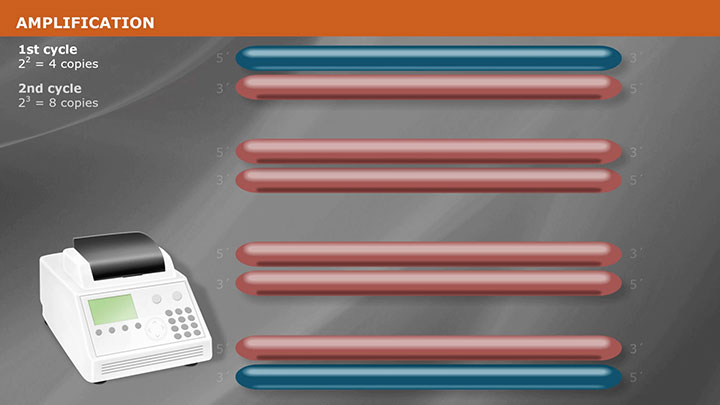

Overview of PCR Cloning




Help us celebrate our 50th anniversary! We have hidden 1,000 golden butterflies and are waiting for you to find them. They can be anywhere that you find NEB! Beginning April 15th, be sure to visit our website and tables at tradeshows and events you are attending. Visit our social media channels frequently for tips on where we have hidden the butterflies – and once you find one, either click or scan the code to be eligible for a 50th anniversary prize pack, as well as a grand prize trip to NEB headquarters in Ipswich, MA!.

To save your cart and view previous orders, sign in to your NEB account. Adding products to your cart without being signed in will result in a loss of your cart when you do sign in or leave the site.