Gibson Assembly Workflow
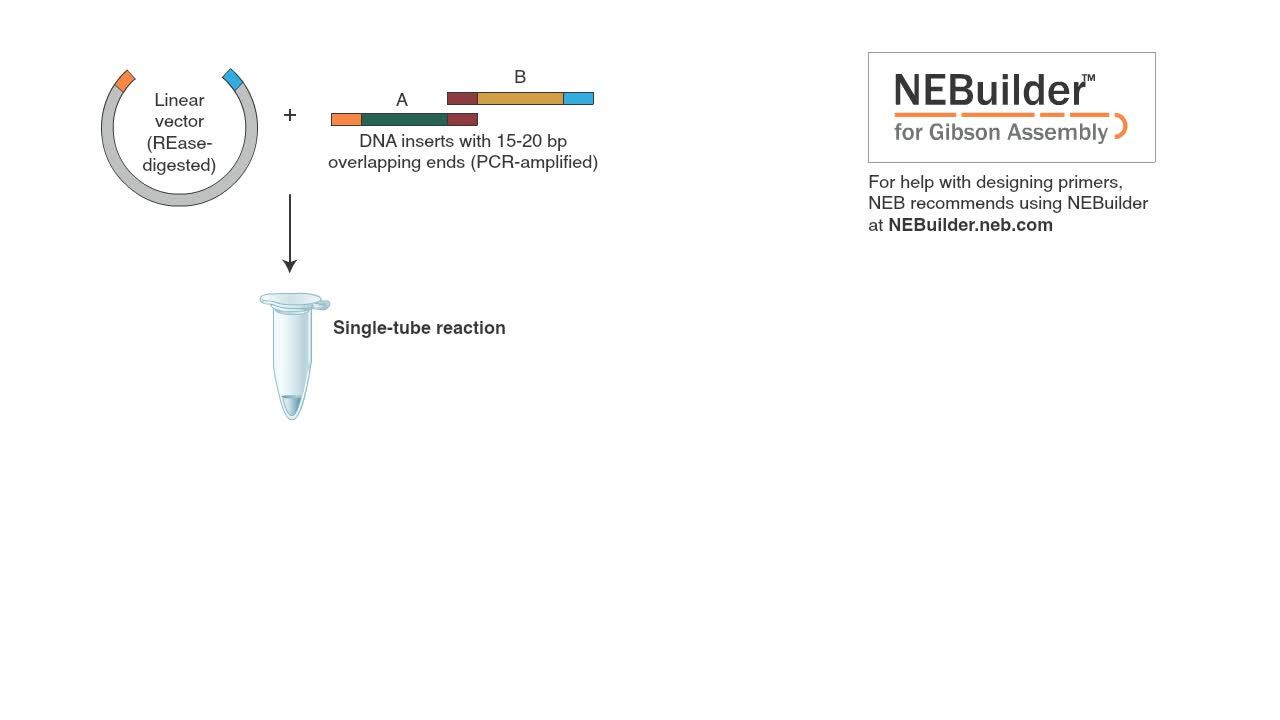
Gibson Assembly™ employs three enzymatic activities in a single-tube reaction: 5´ exonuclease, the 3´ extension activity of a DNA polymerase and DNA ligase activity. The 5´ exonuclease activity chews back the 5´ end sequences and exposes the complementary sequence for annealing. The polymerase activity then fills in the gaps on the annealed regions. A DNA ligase then seals the nick and covalently links the DNA fragments together. The overlapping sequence of adjoining fragments is much longer than those used in Golden Gate Assembly, and therefore results in a higher percentage of correct assemblies. The NEB Gibson Assembly Master Mix (NEB #E2611) and Gibson Assembly Cloning Kit (NEB #E5510S) enable rapid assembly at 50˚C.
Script
Gibson Assembly, developed by Dr. Daniel Gibson, is a robust method for the scarless assembly of multiple DNA fragments in a single tube isothermal reaction.
DNA fragments are designed to have 15 to 20 base pair overlaps that will aid in their proper ordered alignment.
For help with designing primers, NEB recommends using NEBuilder at NEBuilder.NEB.com. The restriction enzyme linearized vector and fragments with overlapping ends are incubated with the Gibson assembly master mix for 15 minutes at 50 degrees Celsius.
In this reaction, three enzyme activities are taking place. Five prime exonuclease, DNA Polymerase, and DNA Ligase. The result is a fully sealed plasmid that is ready for transformation into the included competent cells.
Related Videos
-

Introduction to Gibson Assembly® -
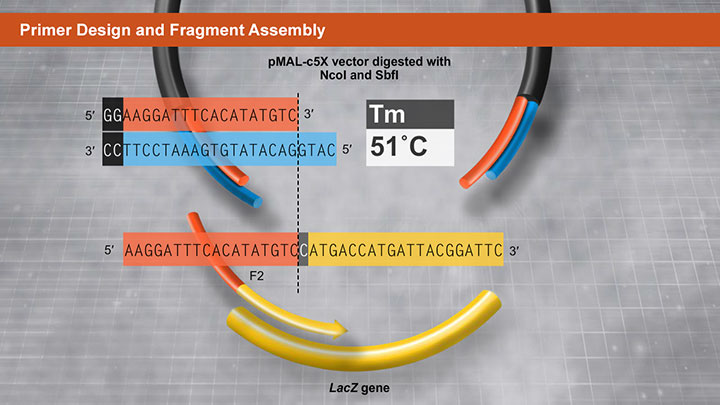
Primer Design and Fragment Assembly Using NEBuilder HiFi DNA Assembly® or Gibson Assembly® -
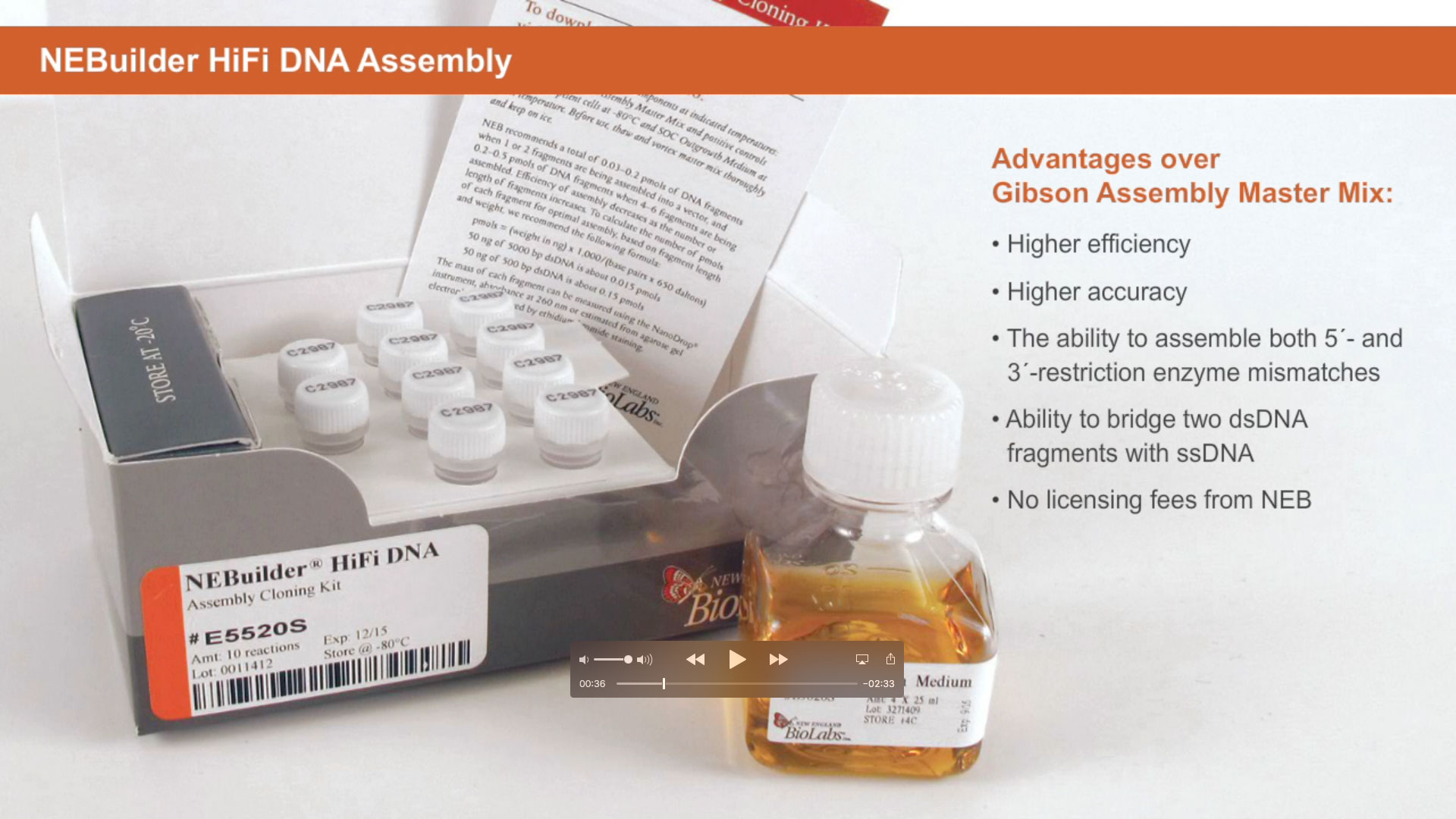
Introduction to NEBUILDER HIFI DNA ASSEMBLY®

