DNA Ligase Fidelity: When does it matter?
Script
Hello everyone. Welcome to today's seminar on DNA Ligase Fidelity. My name is Gregory Lohman. I am a Research Scientist here at New England BioLabs in the DNA Enzymes Division. Today we're going to talk about fidelity. When does it matter, and when is T4 DNA ligase maybe not the ligase of choice?
So, the topics today: we are to do brief introduction into the DNA ligase mechanism. Then talk about T4 DNA ligase and its main applications. Then I would like to discuss DNA ligase fidelity, and what we mean by that, and how that pertains to T4. What applications there are that require high fidelity ligation, and what ligases you might want to use for them. How we here at NEB have profiled DNA ligase fidelity. And then I'll talk briefly about HiFi Taq DNA ligase, a new ligase product that we have targeted towards these high fidelity applications
Okay, so the T4 DNA ligase and other DNA ligases, as you likely know, seal breaks in double-stranded DNA. Illustrated here is a nicked substrate. That's a substrate with a break in one strand of a DNA duplex. And the sealing occurs through the consumption of a co-factor, usually ATP or NAD+. The co-factor is hydrolyzed, and the phosphodiester bond is formed, sealing the break.
This reaction happens in three chemical steps. First, the enzyme reacts independently with its co-factor to adenylate an active site lysine. That adenyl group is transferred from the enzyme to the 5' phosphate at the break. And then that adenylated DNA intermediate is sealed, and the enzyme releases the sealed DNA and the AMP.
The DNA ligase is locked on a variety of DNA structures. The sort of canonical structure, and the one relevant to say, replication, would be sealing a nick, a break in one strand. Ligases are very, very active on nicks. Now, in biotechnology, of course, you're often interested in sealing double stranded breaks. For example, cohesive ends, where you've got a short overlap, complementary overlaps. Blunt ends, where you essentially have fully base-paired ends of DNA that are joined end-to-end. And then the single-base overhang, which is in a sense a cohesive end, but that one base of overlap only makes these, actually, a very poor substrate.
As you go down this list, nicks are very easy to ligate and are the highest activity substrate available. Cohesive ends are between 10 and 100 times slower to ligate than a nick. And blunt ends and single-base overhangs are even slower than that. And so, you'd need a very active ligase to seal these double-stranded breaks, especially the blunt ends.
So this is what you often use T4 ligase for. So here is the activity of T4 ligase on nicked DNA. A few different T4 products. What you're looking at is capillary electrophoresis trace, which we often use at NEB for high-throughput DNA and RNA analysis. It's essentially a gel, separating by size. So, you'll see in the top frame, the nicked substrate unligated with no ligase added, and there's one peak there. After ligation, the peak moves up to a larger size.
And you see here, if we use regular T4 DNA ligase, the quick ligation kit which includes polyethylene glycol, or the Blunt/TA ligation master mix, which is our highest activity formulation of T4. All of them very rapidly seal the nick, essentially to the same endpoint.
Now we look to the cohesive end. Ligase is also very active on cohesive ends and again, all three ligase formulations do very well.
But for these very difficult to ligate fragments, for example a single base overhang. What you'll see is that for T4 in its standard buffer, you only have a very small amount of ligation product. A lot of App DNA, which is that intermediate, that second stage in ligation mechanism. And only in the higher activity formulations including polyethylene glycol do we get really high yields of the difficult to ligate fragment.
Of course T4 DNA ligase is one of the most used ligases in molecular biology and its notable applications include restriction enzyme cloning where you will generally be ligating a short cohesive or blunt end into a vector. And it is also used in modern DNA library adaptor ligation reactions, such as preparing DNA libraries for next-generation sequencing.
Here, you're generally doing a blunt or single-base overhang ligation, a TA ligation. Which requires a very high activity ligase and we usually recommend Blunt/TA master mix or the appropriate NGS library prep kit for those purposes.
The ligases are also used in what we classify as gene synthesis applications. Typically where you're assembling multiple fragments at once. Both applications shown here, Golden Gate and the NEBuilder HiFi Gibson Assemblies can be used to put a single fragment into a vector and often are. But they can also be used to assemble two, six, up to 10 fragments at once. If you'd like to know more about these methods, there's more information on the NEB website. I'm not going to go into too much detail today, except to say that these methods both are dependent on the use of a ligase.
T4 DNA ligase is very high activity. It ligates all sorts of structures, so why not use it for everything? What are its limitations? And when might you not want to use T4?
T4 has high activity, which is why it's so attractive in molecular biology. But with that high activity comes the potential issue of ligation of undesired substrates including, shown here. Gapped substrates on the left of your screen which would be a nick that is missing a base at the ligation junction and T4 can actually ligate across a missing base to make a short loop out. You could also ligate mismatched substrates as shown on the right of the screen there. For example, a substrate with a G-T mismatch.
So, T4's high activity, which is necessary for ligating difficult substrates like blunt ends, could also translate to ligating nicks that are not perfectly base paired or contain gaps. A lot of the time that is not going to be a critical issue but there are some applications where it is.
Let me step back and explain what we mean generally by ligase fidelity. Here we're talking about discrimination against ligating mismatched base pairs. This is sort of discrete from which you might think of as potentially biased if you're ligating two blunt ends together, is there a sequence preference in what ligates?
Here, I'm talking about one strand, the base pairing between strands contains a mismatch. Ligase fidelity has been studied for a number of DNA ligases including T4. And there are some general rules. That is that the higher fidelity ligase, and that higher fidelity on the upstream side of the ligation junction, that is, the base pair providing the 3' hydroxyl for ligation. And weaker fidelity on the downstream side of the ligation junction, which is the base pair providing the 5' phosphate.
Ligases interrogate the shape of the helix. They don't actually check what base pairs are there. What that means is that if a mismatch fits into a normal helix diameter, like GT or TT, it can be favored for mismatch ligation. And, those that form stable, multiple hydrogen bonds, again GT and purine-purines, for example, will tend to pass the fidelity gate of ligases and be ligated. The actual preferences and the degree of mismatch ligation differ from ligase to ligase. And while T4 ligase is fairly promiscuous, there are other ligases that will reject these mismatched base pair substrates and not ligate them.
When do you need to worry about high fidelity in your ligation? We're talking about nick ligation fidelity. You don't need to worry about it for routine cloning when you only have a few overhangs present. If there's only two or three choices of what can ligate and those differ by at least one base pair, it's very unlikely that you're going to insert a short cohesive end in the wrong direction. It's also the case when your goal is to absolutely maximize ligation yield, especially with difficult to ligate structures like blunt ends, then you really, really need a high active ligase like T4. Things like blunt ends and single base overhangs simply don't present opportunity for a mismatch pairing to occur. So, here, again, fidelity isn't important and the need for high activity wins out.
There are more specialized applications where you might need to worry about this fidelity. So, any case when you're going to be ligating a nick and you want to make sure that the nick that you ligate has only correctly base paired bases around the ligation junction, that's when you have to worry about fidelity.
We want to think about applications where you need to specifically ligate nicks without associated end-joining reactions or gap ligation. Often detection of SNPs, single nucleotide polymorphisms through ligation-dependent methods are the really critical applications for high fidelity nick ligation.
And then, finally, cases where you want error-free gene synthesis with multiple fragments being joined together and those are being joined with long overlaps. For example, NEBuilder HiFi takes advantage of high fidelity ligase among other changes to conditions to ensure high fidelity gene synthesis.
So, what am I talking about now when I mean high fidelity ligase? Obviously, hopefully by this point, you understand not T4. But the key high fidelity ligases we sell are the thermostable DNA ligases. Those include Taq DNA ligase, 9 Degrees North DNA ligase, and a recently released new product, HiFi Taq DNA ligase, which is the highest fidelity DNA ligase that we sell right now.
Let me tell you a little bit more about applications that require high fidelity ligation. The sort of grandfather of them all is the ligase detection reaction. The ligase detection reaction is illustrated here. Generally this is a case where you're looking for a specific base pair in a target DNA in some other gene of interest. You will generally then create two LDR probes, which are complementary to the target base pair of interest. As I mentioned earlier, the highest fidelity for most ligases is at the base providing the 3' hydroxyl, so that's generally where you put the probe base that will interrogate the target base pair of interest.
You anneal the probe to your target and ligate them, and high fidelity ligase will only give you a ligation product in the presence of the base pair of interest and not in any variant of bases at that position.
There's the related ligase chain reaction. One of the limitations of ligation detection reaction is that it is a linear amplification. You can only ligate once per target per cycle. The ligase chain reaction attempts to expand this ligation detection to an exponential amplification by using two pairs of probes. One pair that is complementary to one strand of the target, and a second pair that's complementary to the other strand. Generally, you offset these probes so that both probe sets are interrogating the base of interest with the 3' hydroxyl base. But what happens now is when you ligate one set of probes, say, the red set of probes. Those ligated probes now become a template for the yellow probes to anneal and ligate in subsequent rounds. Here, every ligation event generates a new template and you can amplify exponentially.
Ligase chain reaction is known to suffer from high background as any off-target ligation will start to generate more template for further ligation. This is a problem that's been addressed in a variant of LCR called gap-LCR which incorporates a high fidelity polymerase as well as a high fidelity ligase and your probes anneal with a single base pair gap, which a high fidelity ligase will not ligate. That's filled in first by the polymerase and then the HiFi ligase will ligate the filled in probe set.
Some of the more modern methods are the padlock and molecular inversion probes, which are very similar but use a single probe that circularizes on the target. Again, there is both a simply ligation dependent and a gap filling ligation variant of these protocols. And then, once you've created these circles they can be detected by a variety of methods including standard PCR and rolling circle amplification.
Here at NEB, we wanted to first learn about DNA ligases in a very comprehensive manner. A lot of ligation fidelity studies that have been done in the literature focus on one substrate at a time and we were looking for a method that would allow you to sort of get a fidelity fingerprint and look at a large multiplex full of substrates at once to get a picture of how high fidelity a ligase was, and what mismatches were ligated by a specific ligase.
To do that, we devised this system for capillary electrophoresis as illustrated here. Where we would set up a number of targets. One target strand per pool, listed here is the complements. Each complement had a specific pair of bases at the ligation junction. And then we mix with the complement a mixture of eight probes total. Four upstream probes and four downstream probes, each providing all four possible bases on either side of the ligation junction and coded by size.
What that gives us by CE analysis shown here is in the top panel, A. You see the starting probe mixture where the visualized labeled probes, the four phosphorylated downstream probes, run at discrete sizes. And all the ligation products as both the upstream and downstream probes have unique lengths. The ligation products give you 16 products of unique length, all resolvable by CE. Then, in panel C, on the bottom, shows an example of a reaction of that probe mixture with one target strand, the target containing a GT at the ligation junction. And then we can pick out which probe pairs ligate against that particular split. What we see in this case is the CA product, which would be the Watson-Crick product in this case, correctly base paired on both sides of the ligation junction, is the dominant product. We're able to pick out several other products that formed from, in this case, one mismatch each, and are able to identify what those mismatched bases were.
In order to sort of quickly analyze this data we abstracted this into a dot notation. So this is the trace from panel C on the previous page. You see what we do is we encode which peaks we see by position, to identify which product it was. And then we use color to say how high was the yield. Where a green dot would be the highest yield, generally only possible if you're only ligating the Watson-Crick base pair and have almost no mismatch ligation. You see a color code at the bottom. Yellow means significant yield. Red means a pretty large peak but minor compared to the yellow or green. Gray means a peak sort of barely detected.
Then, as we said, we have 16 possible products per target. If we're looking at all combinations of bases at the ligation junction at a target, that gives you 16 possible targets. So, we assemble all of these reactions into a single grid so we can view all products at once. And when looking at these, the Watson-Crick products, correctly base paired on both sides of the ligation junction, will be on the diagonal, indicated here.
Downstream mismatches, that's the 5' phosphorylated base pair are at the positions indicated here, sort of right off the diagonal. The upstream mismatches are these diagonals shown here off of the main diagonal. And then this is them all together where you can see where the Watson-Crick products will run, the downstream mismatches and the upstream mismatches. And all the open squares represent products that are mismatched on both sides of the ligation junction.
With this method in hand, we use it to profile a number of high fidelity ligases available. And I'll note what we do here is we used the manufacturer recommended amount of ligase and supplied buffer. We picked a typical reaction temperature based on the recommendations. In this case, 55 degrees. Though we did do an extended incubation in an attempt to visualize any minor products that would form.
What we see is both Ampligase® and NEB's own Taq ligase do show quite a few mismatch ligation products under these conditions. Both are primarily ligating the Watson-Crick products along the diagonal, but we see substantial ligation of, especially the 5' downstream products. And a few of the upstream products which primarily represent GT mismatches going in the upstream position. As I mentioned earlier, GT is a base pair that ligases have a great deal of difficulty distinguishing from a correct Watson-Crick base pair.
Also shown here is 9 Degrees North DNA ligase, which is a ligase from a different organism, an ATP-dependent ligase or ATP-ligase, attack ligase, or NAD-dependent. And you can see sort of broadly is a little higher fidelity, but what I find most interesting here is that exactly which mismatches are ligated are notably different between Taq ligase and 9 Degrees North ligase. So you can see, for example, a very distinctive pattern for 9 Degrees North where it has those red dots where it really enjoys ligating the sort of same mismatches regardless of what the template is. You see a completely different pattern over on the Taq ligase.
We think this is really good information on its own because it tells you if you need to detect a specific base pair, you can choose a ligase that is discriminatory for the base pair that you're looking for. Now, I also mentioned, of course, quite extensively, that T4 is a low fidelity ligase. So, for comparison, let's see what that looks like.
So, you see with T4, T4 essentially has no ability to discriminate at the 5' side of the junction. All those squares along the main diagonal say it is ligating a mismatched base pair with exactly the same efficiency as the Watson-Crick base pair. And it also does a lot of the upstream mismatches with high activity and a number of even double mismatches were ligated by T4. On top of that, what I'm showing you here is a very low concentration of T4 DNA ligase, 2nM. In a typical sort of cloning reaction, where your goal is to do sticky ends, then blunt ends, you have about 200 times as much ligase present as I have here for this nick ligation. So, T4 simply can't do high fidelity nick ligation. And is not the right ligase to use if that is what you're trying to accomplish.
So, as I mentioned, the high fidelity ligases still show quite a few mismatches. And while you could pick and choose, we were hoping that we could develop, using the screen, a ligase that still gives very high activity on the Watson-Crick product but really knocks down all those mismatch products. You can find out more about the process we used to sort of develop a new buffer. We also looked at different... screened different ligases in all the buffer conditions and the result of that research program is the HiFi Taq DNA ligase shown.
And here you can see compared to Taq ligase that is much higher fidelity. There are only a few things that it still was able to ligate in significant amounts and a few more in trace amounts. But it's greatly improved compared to Taq ligase and shows essentially no 3' upstream mismatches, and only a few downstream 5' mismatches.
We tested out this ligase in a ligase detection reaction application. Here in this application, we ligate one set of probes and then the amount of probes ligated is then quantified by qPCR. So, we can detect a very small amount of ligation product. What you need to know here is that we're looking at the difference in signal between ligating probes on the correct target where everything is Watson-Crick base paired and ligating probes on a mismatched target. So, a higher ∆ Cq means higher fidelity.
What you can see is that both on the 3' mismatch and the 5' mismatch, HiFi Taq DNA ligase beats everything else available. Shown there in yellow is what happens if you put Taq ligase in the HiFi buffer. So, the buffer itself does give some improvement on its own, but the buffer in combination with the new ligase gives the best performance in these SNP detection type applications.
As a happy coincidence, though we were not optimizing for this, we also found the new ligase is significantly more thermostable. It survives with full activity for over 100 cycles where Ampligase™, which is commonly used in cycling ligation reactions, starts to lose activity after 25 cycles and has only a small amount of activity left after 100.
I do want to mention one more thing. We talked about how ligase affects fidelity, and I mentioned optimizing buffer helps fidelity, but that's not the only consideration when doing these applications. Temperature can affect both yield and fidelity and it is going to be temperature as it relates to probe annealing temperature. So, if you're too far below the annealing temperature, the probes shown here in the 45 degree side, the amount of mismatch ligation goes up dramatically. And you can end up, even for a high fidelity ligase, shown here is standard Taq DNA ligase. A loss of preference, especially at the 5' side of the ligation junction.
When you're closer to the annealing temperature, shown here in the middle of 55, you get a much improved yield of the Watson-Crick products and a suppressed yield of the mismatch products. Now, if you go even higher, well above the Tm, you do knock down the mismatch products, but you also start to lose yield of your desired product because here, simply, the probes aren't properly annealed.
In order to really maximize fidelity in an application like this, you need to both use a high fidelity ligase and choose your reaction temperature appropriate for your probe set and the buffer you're using. To help you do that, we recently released a Thermostable Ligase reaction temperature tool shown here and available at the URL shown at the bottom of the screen.
We really recommend, if you're designing any of these detection ligation-dependent, detection reactions, or in general going to be using a high fidelity ligase to ligate probes, you're going to get the best specificity if you use this tool to optimize your probe design and then choose the correct ligation temperature for your probe set.
I just want to summarize today. We talked about, of course, T4 ligase being very high activity in end-joining ligation, but it can become a problem is your goal is to ligate nicks and that T4 will readily ligate many mismatches in nicks. The high fidelity is not needed in routine cloning applications but it is critical in SNP detection reactions. NEB devised a screen for rapid profiling nick-ligation fidelity and this was applied in the development of HiFi Taq DNA ligase, which we now recommend as the highest activity and the highest nick-ligation fidelity available commercially for use in SNP-detection applications and related technologies.
We have a few references on today's talk. A few references there at the top about the overview of ligation and ligation chemistry. In the middle section, we have ligase fidelity references, including the third one there is the overview of our mismatch detection profile. And then, the bottom screen there we have some information for you on high fidelity ligation applications and a link to the HiFi Taq DNA ligase product page.
Many people participated in this work at NEB and everything shown here is dependent on the great group of collaborators I had available internally and externally. Just a few people, including my boss Dr. Tom Evans. My collaborators Jennifer Ong, Vladimir Potapov, Nicole Nichols, Rob Bauer, Tom Jurkiw, Luke Sprenkle, and Christina Manner have all contributed very much to this ligation research.
We also have some application development specialists: Eric Cantor, Ashley Luck, Mary Lorenzen and Bo Wu. Who, between them have developed and maintained the ligase products.
Everything we do here is dependent on the NEB sequencing core and the staff listed there. As well as the founders and directors of NEB. And, I'd also like to thank NEB marketing and Marcomm for helping me set up this seminar and delivering it to you. And, my collaborators externally who are just great resources to have. And it's a really special thing to work at NEB where you can work in an industry on these very interesting research problems. And I've just had great feedback and back and forth with academic researchers in your area.
And now, I'd like to open it up for any questions. Thank you very much.
Okay, so, now I'm going to try and address some of these questions. Thank you all for listening. All right, let's see, what do we got. I'm going to do some easy ones first.
So, have we looked at untemplated ligation? Yes, I have. We've actually looked at both DNA templated, RNA templated and untemplated ligation for most of the ligases that we sell. Untemplated DNA-DNA ligation is an extremely difficult reaction and very few things catalyze it. So, T4 DNA ligase has essentially no detectable activity on untemplated DNA to DNA ligation. Some of the RNA ligases obviously are RNA-RNA single stranded specific, or will at least do that reaction. And some of those RNA ligases will do RNA to DNA with decent efficiency, but DNA to DNA is a very difficult reaction.
Okay, so, let me see. Any applications where it is an advantage to seal across a gap? That's actually an interesting one. As I mentioned, there's these gap ligase variants of the LCR and the molecular inversion probes where you're using a gap as sort of a stop and then filling it in order to ensure high fidelity. But to actually, intentionally ligate across a gap, I'm not offhand aware of any applications. But, yeah, it's potentially interesting. There certainly could be, but once you start to get longer insertions, like if you're talking about more than one or two nucleotide gaps, I don't think a lot of ligases will do that efficiently because... certainly not the double stranded DNA ligases because they kind of get... the loop out gets in the way of proper binding to the active site.
Okay, what would we recommend for ligating hairpin adapters onto template DNA with random sticky ends. This is actually an area of active research for my group, what we've moved onto now from the nick ligation is trying to profile the fidelity of sticky end ligation. And to profile the sort of sequence bias of blunt ligation.
So, for sticky end ligation, so we're not ready to share the details of that but I can give some general rules there. The short version is that the bad actor again is GT mismatches, even in a sticky end situation under some case the ligase has trouble distinguishing that from a correctly templated base pair. We still generally recommend T4 because the cohesive end ligation with short ends, talking about like five bases or less, the thermostable high fidelity ligases have almost no activity on those substrates. So, you need T4, T7 also works as does T3 DNA ligase. But our preference is for T4 because of high yield and high efficiency. I would use for that the regular standard T4, don't use either of the products containing polyethylene glycol in the buffer because that does modestly increase mismatches for the short sticky ends.
And then I actually, in terms of temperature, would recommend doing that at room temperature or even up to 37. Traditionally people like to do four degrees, 16 degrees overnight, I actually don't think that's necessary at all, which is somewhat of a heretical thing to say, I know. But I find you get really good yields at room temperature or even elevated temperature because despite the fact that the sticky ends are predicted to be annealed, T4's activity is so high at those temperature that it sort of grabs them and ligates them as they anneal. And you can really suppress those mismatches because a single mismatch makes a huge difference at 25 to 37. So, that's what I would recommend for that.
What else we got? So, how does T7 compare with T4? That's an interesting one. I don't have a comprehensive answer on that yet. Again, that's something I think we will have. The short version there is that some people prefer T7 for high fidelity sticky ends. We know that T7 doesn't really do blunt ligation, so it only does cohesive end ligation. So, there's that interesting difference if you want to do cohesive ends but not blunts, you can use T7. I'm unconvinced that T7's sticky end fidelity is actually that much better than T4, so I'd still recommend using T4 in its standard buffer. But, you know, there are many applications with people out there who use it. Some people who do Gibson prefer T7 to T4, oh, excuse me, not Gibson, Golden Gate assembly, prefer T7 to T4. So, I would just recommend seeing what's been published and it's potentially worth comparing the two in your own application. But my default recommendation remains T4.
Kinetic information. Yeah, so I've actually published a lot on DNA ligase kinetics. If you search for papers I published in JBC, NAR and PLoS One, my group actually does a lot on just the basic kinetics, particularly with T4 DNA ligase. So, we have published those numbers. The approximate turnover rate of a ligase on nicks is about one per second and the KM is extremely tight, like single digit nanomolar KM. But it differs from ligase to ligase. Getting KM and turnover rate for double stranded break fragments is trickier so we don't have firm numbers on those, but the KM for blunts is roughly in the high micromolar range, so it's much weaker binding than binding to a nick.
Does Taq ligase have any activity on RNA-splinted DNA molecules? No, it does not. None of the high fidelity thermostable ligases accept hybrid helices with the exception of, they will take... if the RNA is providing the hydroxyl to the ligation, so if it's that upstream probe position and everything else is DNA... Taq, 9 Degrees North will do those. HiFi Taq will not. HiFi Taq is fully selective for fully DNA substrates. For RNA-splinted ligation, the best ligase we have for doing that is SplintR ligase, which is unfortunately not thermo-stable. That's SplintR if you search that in our catalog. That is about a thousand times more efficient than T4 DNA ligase for RNA splinted DNA. And then both T4 RNA ligase 2 and DNA ligase will do RNA splinted DNA with modest activity, but we recommend SplintR ligase for that.
Let's see. The temperature data shown there was shown for standard Taq ligase. T4 DNA ligase is rapidly inactivated above 45 degrees, and so 37 is the highest temperature I recommend using T4. And even there, you're just not going to get the... the fidelity data I showed for T4 was at 37 degrees and that's essentially as good as it gets for nicks. Again, if you're going to do high fidelity sticky end ligation, 30 to 37 is a good temperature to try for T4. The thermostable ligases are active 37 and above. And they're the ones you want to key to the Tm of your probes or the nick that you're trying to ligate.
HiFi ligase will only work on fully DNA structures, it won't tolerate RNA in any position. Okay, interesting question we often get. Is there any way of terminating the activity of any of these thermostable ligases, say, Taq or 9 Degrees North after ligation. So, they're all resistant to cycling as you see, but sometimes, if you want the activity cleared after you've run the reaction there's really not a lot you can do except purify the DNA. The only other... the way to kill the activity is to add either SDS or EDTA but of course that's going to interfere with most downstream reactions you're going to want to do. So, cleaning up the DNA is really the only option to get rid of the ligase activity.
Now, that said, we have not...in a lot of follow on reactions, the residual ligase activity is not going to be a problem, especially if next steps are at lower temperature. Or if the next steps are not going to be generating nicks, that left over ligase activity won't really cause issues. So, for the qPCR-coupled LDR that I showed a little bit of data from and there's some more about that on our website for the HiFi ligase. You'll find what we did there was we did a ligation and then carried an aliquot of ligation mixture straight into the qPCR reaction. And the ligase was still there but it didn't interfere with the downstream steps in any way that we could see.
What else? Just give me a second to review the questions remaining. Question on length of the strand. So, ligases prefer about 10 base pairs on either side of ligation junction. Activity starts to drop off the shorter they get. Longer doesn't seem to have a strong preference, though if you're interested... once you start to have a lot of DNA around, for example, if you have just a few ligatable ends and a huge mass of genomic DNA, or you have very, very long substrates, like, you know you're trying to put a 10,000 kb insert into a vector or something, you might notice inhibition in the ligation speed.
We actually published on this recently in PLoS One on T4. But ligases seem like they're searching DNA by binding randomly. So, a very large amount of non-nicked DNA can end up inhibiting the ligation reaction, simply from like a search problem. The ligase can't find its target. But you don't get... we have not yet found a place beyond... once you get beyond that minimum binding site length of about 10 bases on either side, you don't really see a huge increase. It's not like it prefers 30-mers or 40-mers or 50-mers. It's pretty much flat until the amount of DNA present in the reaction starts inhibiting things.
I have a question on multiplexing SNP detection. We have not done it. We have not done a multiplex LDR reaction, though, sort of conceptionally, the way we do the screens are multiplexed, where we've got a whole bunch of probes competing for a single target. We have not multiplexed multiple probes, multiple targets in the same reaction. I don't see any reason why you couldn't. It should work if you have a reaction designed for Taq ligase or Ampligase®, HiFi ligase is designed to be a drop-in into those reactions, that you should be able to. Even if... you can use a different buffer if you would prefer, although you'll get the best fidelity with using both HiFi ligase and HiFi buffer but you should be able to use our new buffer ligase system essentially comparably in any protocols you're currently using Taq or ampligase for SNP detection.
Answer a question. Do we have any idea why HiFi Taq is such a high fidelity? Indeed we do. Unfortunately, I cannot share why it does. You can look at... if you look at our NAR paper on the development of the assay system we do describe what is essentially the HiFi ligase buffer development and in there the really key component is higher pH, which gives both higher activity and seems to give a bit more selectivity. And the really important thing is salt. At higher salt there is an effect on kcat/KM, that's probably a binding effect. Where the binding of both to fully base paired substrates and mismatched substrates is weakened, but the mismatch is weakened more. So you separate the relative kcat/KMs of those two, and under those typical conditions you still get really efficient fully base paired ligation, but you've wiped out binding of mismatches unless you don't ligate those very efficiently.
So that's a buffer component. The ligase component is proprietary.
Yes, there are, so okay, a question on the assay. Do we use specific sequences? Yes. You can get all those details in the Nucleic Acids Research report on that study which is available online.
Question on annealing adapters, sort of annealing hairpin adapters. So, if you're talking about blunt or TA adapters, annealing those doesn't do anything for you. You can't meaningfully anneal those. For longer overhangs, like if you're thinking about something like a four base overhang system, comparable to a Golden Gate assembly, there's debate about whether annealing matters even between me and my colleagues here. I don't think so. I generally prefer doing those kind of reactions, just running them at room temperature and you're basically dynamically annealing at that point. And T4 will grab and react things as they anneal. And you're not going to gain any huge benefit from cycling.
Now, if you look at our Golden Gate assembly protocol, that kit does give two versions of the protocol, one for continuous incubation at 37 and one for a cycle protocol. And there is some evidence that for really complex assemblies, cycling helps. And that may get around cases where you've got really stably annealed mismatched substrates, ones that won't ligate, but you can remix if you melt. But in my experience, if you just incubate at 37, most short overhangs are going to scramble over the course of the reaction.
Just a few more questions, I think, and then we'll wrap up here. So, if you've got anything else get it in now.
Can ligase form chemistry multiple times in a single binding event? Yeah. So if you go back into some of our kinetics papers on T4 which we did largely on nicks. You'll find that ligase will turn over on nicks thousands and thousands of times. I've similarly found for T4 at least, it does, indeed turn over on blunt ends and cohesive ends. For blunt ends the reaction is slow, so you generally swamp the reaction with more ligase than there are ends to get the reaction done in a timely fashion. But it will turn over albeit very slowly, on blunt end substrates.
Other ligases, I have not characterized as well. Everything I've looked at turns over efficiently on nicks, but I can't say for sure on end-joining reactions.
And we have a question on template purity. How does template purity affect DNA ligation? Actually good, this is a good question. One random point I was actually asked to make. We get a lot of questions at NEB that come down to my cloning had trouble, my library prep had trouble, and people often like to blame the ligase. Usually it's not the ligase that causes the ligation step to fail. It tends to be DNA quality. So, there are a lot of issues. DNA ligases are sensitive to salt. So, if you're trying to do a cell lysate with a whole bunch of salt and potentially a whole bunch of detergent....ligases are not going to work in that kind of mixture.
If you need to do a ligation in high salt we recommend T3 DNA ligase, but even there, the salt tolerance tops out around 250 millimolar. If you have a whole bunch of DNA around, again, as I mentioned, that's going to... the ligase is going to waste its time searching all that DNA. So, if you've only got five ends per a four gigabase genome you're trying to ligate, ligases will have a lot of trouble with that. And then, generally low quality DNA. You know, DNA that has a lot of gaps, and has a lot of frayed ends. Ligase is going to have a lot of trouble with that. You know, T4 can ligate across gaps and ligase mismatches, but it's got reduced activity on those substrates. So, if you're trying to do an efficient ligation in DNA that's badly damaged, that hasn't been end repaired. That hasn't been treated with DNA repair, and hasn't been cleaned up by column purification or beads or anything like that, yeah, that will definitely impact your ligation efficiency.
All right. Thanks, everyone for attending our DNA ligase fidelity seminar. Thanks very much for the great questions. I just want to let you know that we will have a recording available. Anyone who's registered will get an email in the next little while with details on how to access that recording. Thanks again, and we hope to see you at future webinars.
Related Videos
-

DNA Ligation -
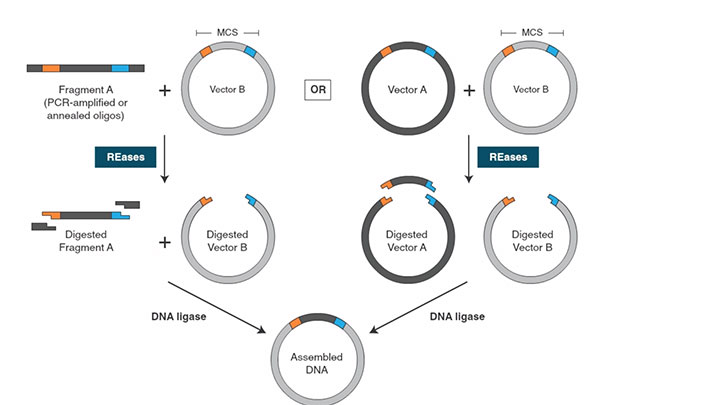
Traditional Cloning Workflow -

What are the best conditions for DNA ligation?

