Construction of an sgRNA-Cas9 Expression Vector via an ssOligo Bridge
Script
NEB has developed a protocol to quickly and easily change the targeting of a single-guide RNA in a sgRNA plasmid or to create an sgRNA plasmid library. The method uses a single DNA oligonucleotide, a restriction enzyme digested plasmid and the NEBuilder HiFi DNA Assembly Master Mix.
Choose a target sequence or target sequence using a design tool of your choice. Design a single-stranded DNA oligo, containing a target sequence flanked by a partial promoter sequence and scaffold RNA sequence. In this animation the U6 promoter is used.
Prepare the DNA oligo. Assemble the reaction mix by combining DNA oligo, restriction-enzyme linearized plasmid and water.
Add NEBuilder HiFi DNA Assembly Master Mix and incubate for 1 hour at 50 degrees Celsius.
Transform NEB 10-beta Competent E.coli with the assembled product.
Spread outgrowth on plate with antibiotic and incubate overnight at 37 degrees Celsius.
Pick colonies to grow, and purify the plasmid DNA for sequencing.
Some traditional methods require synthesis, phopsphorylation, annealing, and ligation of two oligos into a digested and dephosphorylated vector. This new protocol using a single DNA oligonucleotide with NEBuilder HiFi DNA Assembly Master mix is a simple and streamlined way to rapidly create specifically targeted Cas9/sgRNA plasmids.
Related Videos
-

NEB® TV Ep. 12 – Applications of DNA Assembly -

NEBUILDER HIFI DNA ASSEMBLY® Removal of 3´ end Mismatches -
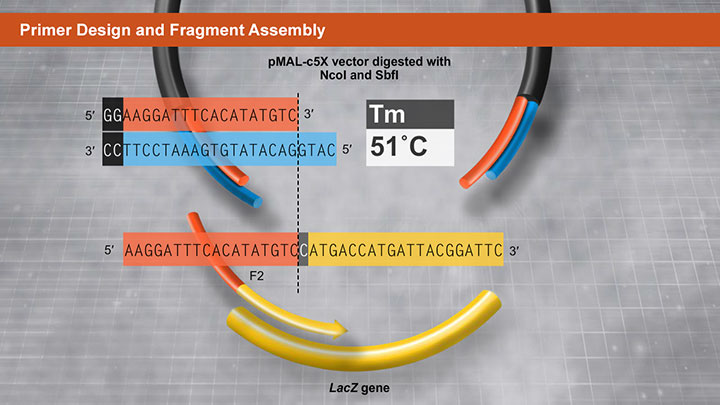
Primer Design and Fragment Assembly Using NEBuilder HiFi DNA Assembly® or Gibson Assembly®

