Hydroxymethylation Detection and Analysis
Hydroxymethylation is a highly dynamic, low abundance DNA modification with considerable potential as an epigenetic biomarker for development and disease states. Mammalian DNA is enzymatically modified at the 5th carbon position of cytosine (C) bases to 5mC, predominantly in the context of palindromic cytosine-guanine (CpG) dinucleotides. 5-Methylcytosine is amenable to enzymatic oxidation to 5-hydroxymethylcytosine (5hmC) by the TET family of enzymes.
Hydroxymethylation has distinct functionality apart from methylation, but a complete understanding of 5hmC has been obscured by legacy DNA methylation detection methods that do not discriminate 5hmC from 5mC. Bisulfite conversion is a harsh chemical treatment that deaminates unmodified cytosines to uracil, without deaminating the 5mC and 5hmC modified bases. The treatment damages DNA leading to biased genome coverage. Bisulfite conversion can be followed by PCR amplification, or massively parallel sequencing methods to reveal cytosine methylation status in specific genes or whole genomes. However, both 5mC and 5hmC readout as cytosines, therefore PCR and standard bisulfite sequencing methods are unable to distinguish between 5mC and 5hmC. Other base-conversion techniques and enrichment methods designed to discriminate 5hmC require higher depth of sequencing or higher sample inputs. Sensitive and specific detection methods are required to elucidate this low-abundant modification's functionality.

5hmC Kits and Reagents
The NEBNext® Enzymatic 5hmC-seq conversion module (NEB #E3365) is a robust method to directly detect 5hmC at single-base resolution from 0.1 – 200 ng of DNA. The two-step enzymatic conversion workflow generates high yields and even GC coverage. After fragmentation, end repair and dA-tailing and ligation to sequencing adaptors, 5hmC in DNA libraries are glucosylated by T4 β-glucosyltransferase (T4-βGT), protecting it from deamination. In the next step, APOBEC deaminates 5mC to thymine and unmodified cytosine to uracil, while the protected 5hmC is not converted. The NEBNext Enzymatic 5hmC-seq Kit (NEB #E3350) includes library prep reagents for Illumina® sequencing workflows. E5hmC-seq data can be combined with enzymatic methyl sequencing (EM-seq) to detect and differentiate 5mC and 5hmC. NEBNext Primers for Epigenetics (Unique Dual Index Set 2B) (NEB #E3392) enables multiplexing with the NEBNext E5hmC-seq™ Kit. Critically, these methods have low input requirements and are compatible with clinically relevant sample types such as cell free DNA (cfDNA).
Liquid Chromatography Mass Spectrometry (LC-MS) also allows for high resolution detection of 5hmC. The Nucleoside Digestion Mix (NEB #M0649) is an optimized enzyme mixture that digests epigenetically modified (m5C, hm5C, f5C, ca5C, m4C, m6A, etc.), unnatural, or damaged bases to generate single nucleotides from DNA in a one-step method to prepare for LC-MS.
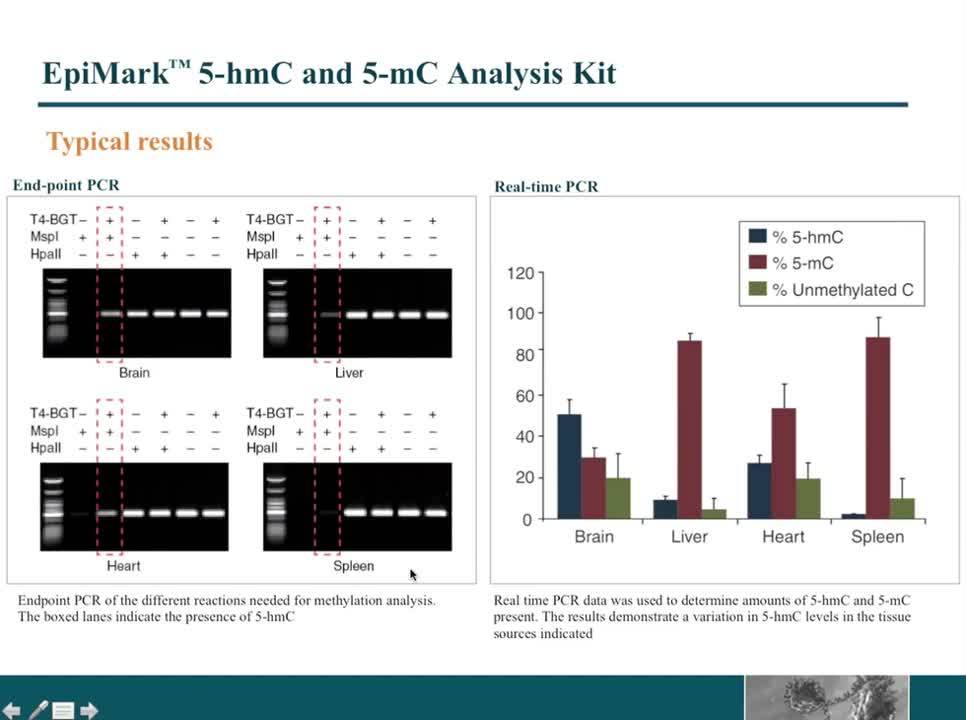
Quantitative PCR analysis will demonstrate the relative abundance of hydroxymethylation when DNA is treated with T4 Phage β-glucosyltransferase (T4-BGT) (NEB #M0357) and then digested with methylation-sensitive restriction enzymes. The covalent attachment of glucose by T4-BGT results in differential substrate specificity for 5mC versus 5hmC. When 5hmC occurs at the internal CG of CCGG, this glucosylation of internal CG blocks the recognition site for the methylation insensitive restriction enzyme MspI (NEB #R0106). HpaII (NEB #R0171) is an isoschizomer of MspI that is blocked by CpG methylation. Quantitative PCR based methods can be used to determine relative abundance of C, 5mC and 5hmC at the internal CpG site of CCGG sequence.
- NEBNext Enzymatic 5hmC-seq Kit (NEB #E3350) enables direct detection of 5hmC using NEBNext Ultra™ II library prep reagents, a two-step enzymatic conversion workflow, and the E5hmC-seq Adaptor optimized for Illumina Sequencing.
- NEBNext Enzymatic 5hmC-seq Conversion Module (NEB #E3365) enables enzymatic conversion for detection of 5hmC from 0.1 – 200 ng of DNA.
- NEBNext Primers for Epigenetics (Unique Dual Index Set 2B) (NEB #E3392) and NEBNext Primers for Epigenetics (Unique Dual Index Set 3) (NEB #E3404) enable higher levels of multiplexing with minimized index hopping with the NEBNext E5hmC-seq™ Kit. The sets are compatible for combination to multiplex up to 120 reactions.
- Nucleoside Digestion Mix (NEB #M0649) digests epigenetically modified bases to generate single nucleotides from DNA in a one-step method to prepare for LC-MS.
- T4 Phage β-glucosyltransferase (T4-BGT) (NEB #M0357) glucosylates 5hmC only.
- HpaII (NEB #R0171) cleaves unmethylated internal CpG sites.
- MspI (NEB #R0106) cleaves methylated internal CpG sites
Illumina® is a registered trademark of Illumina, Inc.
Choose Product:
Choose Type:
- What types of epigenetic modifications can be identified using the Nucleoside Digestion Mix?
- Do E5hmC-seq™ libraries detect 5mC?
- How deep should E5hmC-seq™ libraries be sequenced?
- What types of samples can be processed using the NEBNext® Enzymatic 5hmC-seq Kit (NEB #E3350) and the NEBNext Enzymatic 5hmC-seq Conversion Module (NEB #E3365)?
- NEBNext Selector
- Comparison of Methods for DNA Methylation Analysis
- Dam-Dcm and CpG Methylation
Web Tools
Selection Tools
Products and content are covered by one or more patents, trademarks and/or copyrights owned or controlled by New England Biolabs, Inc (NEB). The use of trademark symbols does not necessarily indicate that the name is trademarked in the country where it is being read; it indicates where the content was originally developed. All other trademarks are the property of their respective owners. The use of this product may require the buyer to obtain additional third-party intellectual property rights for certain applications. For more information, please email busdev@neb.com.
This product is intended for research purposes only. This product is not intended to be used for therapeutic or diagnostic purposes in humans or animals.
Ineligible item added to cart
Based on your Freezer Program type, you are trying to add a product to your cart that is either not allowed or not allowed with the existing contents of your cart. Please review and update your order accordingly. If you have any questions, please contact Customer Support at freezers@neb.com or 1-800-632-5227 x 8.