What is Cappable-seq?
Script
Cappable-seq
Laurence Etwiller:
Cappable-seq exploits the fact that primary transcripts in prokaryotes have a unique triphosphate at the beginning of the RNA.
Cappable-seq uses this feature to capture the 5′ end of these molecules, enabling determination of transcription start sites in prokaryotes at single base resolution.
In addition to TSS determination, Cappable-seq depletes ribosomal RNA and reduces the complexity of the transcriptome to a single quantifiable tag per transcript enabling digital profiling of gene expression in a microbiome.
Are there other methods to identify transcription start sites?
For prokaryotes, dRNA-seq has been used extensively to identify TSS. It uses the terminator exonuclease (TEX) treatment to theoretically remove all non-triphosphate RNA.
The limitation of this technology is that success seems to highly depend on the 5' end structure of the RNA. Therefore, removal of the non-triphosphate RNA (non-TSS) is incomplete and in some cases non-existent, leading to false positive TSS (ex : tRNA) and large ribosomal contamination.
The advantage of Cappable-seq over the TEX method is the direct capturing of the triphosphate (as opposed to the depletion of the non-triphosphate) which leads to higher specificity, removal of rRNA and improved tolerance for low RNA quality.
What is SMRT-Cappable-seq? What are the key differences and what kinds of biological questions can be studied with this technique?
We recently developed SMRT-Cappable Seq, a derivative of Cappable-seq that sequence full length RNA using PacBio as oppose to Cappable-seq that identify the TSS only.
We applied SMRT-Cappable Seq to the best annotated prokaryotic genome, E.coli, and demonstrated that 34% of operons in E.coli are longer than the known annotations by at least one additional gene(s).
Phasing operons also enabled to observe operon variants and an granularity in the operon structure of E.coli that is reminiscent of eukaryotic splicing. SMRT-Cappable-seq therefore describe complete genome-wide landscape of bacteria isoforms. Similar to capable-seq, SMRT-cappable-seq also significantly remove rRNA.
Related Videos
-
NEB® TV Ep. 21 - RNA modifications -
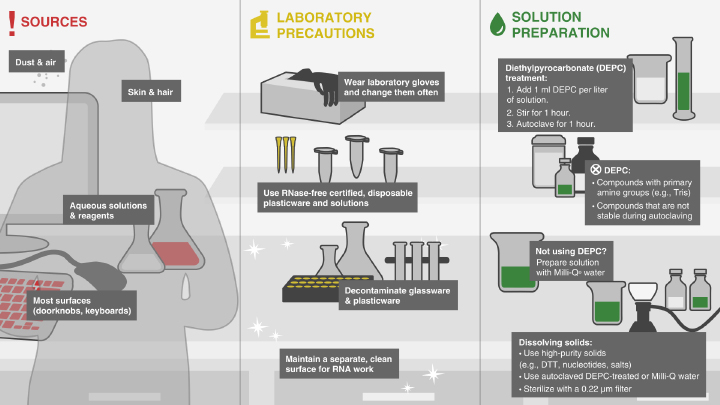
Avoiding RNase Contamination -

In Vitro Transcription and Capping of Gaussia Luciferase mRNA Followed by HeLa Cell Transfection

