High Sensitivity 5-hmC Detection in Balb/C Brain Tissue

Follow NEB Product Development Scientist, Romas Vaisvila, Ph.D., as he demonstrates the EpiMark® 5-mc and 5-hmc Analysis Kit for locus specific identification and quantification of 5-hmc in this protocol "High Sensitivity 5-hmc detection in Balb/C Brain Tissue".
Script
The overall goal of the following experiment is to analyze and quantitate 5-methylcytosine and 5-hydroxymethylcytosine within a specific locus of genomic DNA. This is achieved by glucosylating the hydroxyl group of 5-hydroxymethylcytosine, which converts a cleavable MspI site into a non-cleavable site. As a second step, the genomic DNA is digested by the restriction enzymes, MspI and HpaII, which utilize their differential methylation sensitivities to identify glucosylated 5-hydroxymethylcytosine sites.
Next, PCR is used in order to interrogate the locus. Results are obtained that show the presence of 5-hydroxymethylcytosine in a specific locus, qPCR can be used to quantitate the amount of 5-methylcytosine and 5-hydroxymethylcytosine.
Theodore Davis:
The primary advantage of this technique over other methods, such as bisulfite sequencing, is that you can discriminate between 5-methylcytosine and 5-hydroxymethylcytosine. Performing the experiments today will be Romualdas Vaisvilla. Romualdas was a primary developer of this kit at New England Biolabs.
Anthony Sheets:
Prior to the start of this protocol, purify the genomic DNA from BALB/c brain tissue as described in the written procedure. In a 1.5 milliliter reaction tube, mix the genomic DNA, UDP-glucose, NEBuffer 4, and enough nuclease free water to bring the total volume to 310 microliters. Split this reaction mixture into two tubes of 155 microliters each. Then add 30 units, or three microliters, of T4 beta glucosyl transferase to one tube.
Mix well by gently pipetting up and down. Do not add the T4 BGT to the second tube, as it is the control reaction, but be sure to add three microliters of water. Incubate both tubes at 37 degrees Celsius for 12 to 18 hours, during which time the BGT will add glucose to the hydroxyl group of 5-hydroxymethylcytosine groups in the sample.
To begin the restriction enzyme digestion, aliquot 50 microliters of the reaction mixture into each of three 0.2 milliliter PCR strip tubes, labeled one through three. Then, aliquot 50 microliters of the control mixture into each of three PCR strip tubes, labeled four through six. Add 100 units of MspI into tubes one and four. Next, add 50 units of HpaII into tubes two and five.
And mix well by gently pipetting up and down. Do not add anything to tubes three and six, as they are controls. Incubate all six tubes at 37 degrees Celsius for at least four hours. Then, add one microliter of proteinase K into each tube and incubate at 40 degrees celsius for 30 minutes.
Finally, inactivate the proteinase K by incubating at 95 degrees Celsius for 10 minutes. This procedure is demonstrated using NEBs LongAmp Taq for end-point PCR. However, other PCR reagents and procedures can be substituted. To a 0.2 milliliter PCR reaction tube on ice, add 5X LongAmp Taq reaction, buffer, dNTPs, forward and reverse primers, the previously obtained template DNA, LongAmp Taq DNA polymerase, and enough nucleus free water to achieve the final volume of 50 microliters.
Gently mix the reaction. If necessary, collect all liquid to the bottom of the tube by a quick spin. Transfer the PCR tubes, from ice to a thermo cycler with the block preheated to 94 degrees Celsius, and start the cycling program. For a routine three step PCR, there should be one initial denaturation step at 94 degrees Celsius for 30 seconds, followed by 30 cycles of 94 degrees Celsius denaturation for 15 seconds, 55 to 65 degrees Celsius annealing for 30 seconds, and a 65 degrees Celsius extension for 20 seconds, or 50 seconds per Kb.
This should be followed by one final extension at 65 degrees Celsius for five minutes. Real-time PCR can also be performed to quantitate amounts of 5-methylcytosine and 5-hydroxymethylcytosine. Please refer to the product manual, found on neb.com, for specific information.
A comparison of 5-hydroxymethylcytosine amounts at locus 12 was performed using endpoint PCR and real-time PCR. Four different mouse BALB/c tissue samples were analyzed, including brain, liver, heart and spleen. The variation in 5-hydroxymethylcytosine is apparent in the box to gel lane of the endpoint PCR samples.
For comparative purposes, real-time PCR data was normalized to uncut DNA and a standard curve was used to determine copy number. The samples were normalized by dividing the copy number of samples by the copy number of the undigested control. Here, the variation in 5-hydroxymethylcytosine can be seen by comparing the yellow bars.
Theodore Davis:
A few notes on performing the technique. It's critical that all the enzymatic reactions are carried to completion. For the BGT, it's important that you go overnight, so that you can glucosylate all the DNA sites. For MspI and HpaII, it's important that you go at least four hours with those reactions to make sure that you reach completion. And finally, we provide DNA controls and primer controls, so that you can carry out control reactions side by side.
Related Videos
-
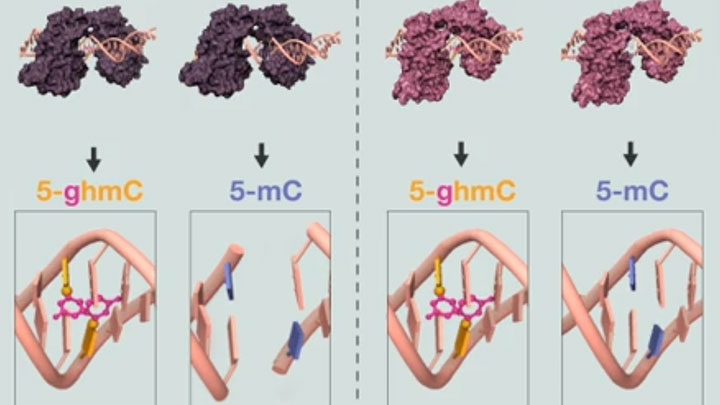
EpiMark 5-hmC and 5-mC Analysis Kit Protocol Animation -
Interactive Tutorial Explaining the Phenomenon of Epigenetics at a Molecular Level -
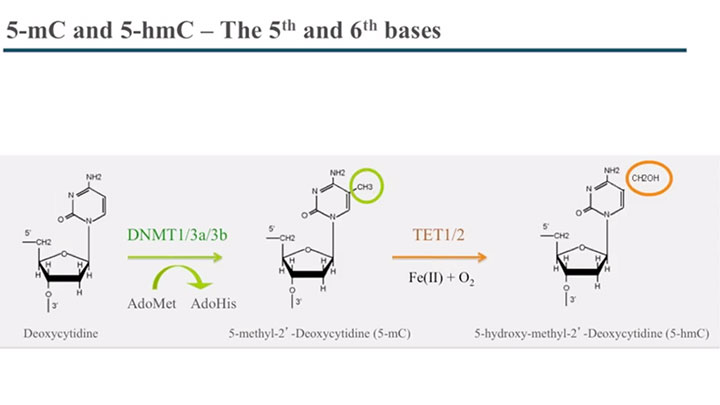
Locus Specific and Global 5-hydroxymethylcytosine Detection and Quantification

