Base modifications affecting RNA polymerase and reverse transcriptase fidelity
Script
Hello, my name is Jennifer Ong, and I'm here to tell you about a recent paper that we published in Nucleic Acids Research.
RNA modifications are biologically important. Modifications, such as pseudouridine, N6-methyladenosine, or 5-methylcytosine, are found in messenger RNA from a variety of organisms, including eukaryotes, where they can alter gene expression or mRNA stability.
RNA modifications are involved in various cellular processes or diseases, and so, there's a lot of interest in detecting and quantifying RNA modifications in the transcriptome especially by next generation sequencing. However, in spite of all this interest, not much is really known about how these modifications affect polymerase fidelity.
We designed a fidelity assay based on PacBio single-molecule real-time sequencing, which is capable of sequencing individual DNA molecules with very high accuracy. In our assay, we synthesized RNA from a pool of nucleoside triphosphates using T7 RNA polymerase. This nucleotide pool can contain standard NTPs, or we can replace one of the bases with a modified base. The unmodified or modified RNA is then reverse transcribed into cDNA, which is then converted into double-stranded DNA and sequenced.
After sequencing, we can then compare the results to the reference to determine an error rate. This first strand error rate is then the sum of the error rates from both T7 RNA polymerase and the reverse transcriptase.
We measure the combined error rates for T7 RNA polymerase and ProtoScript II reverse transcriptase with standard and modified RNA, including N6-methyladenosine, pseudouridine, 5-methylcytosine, 5-methyluridine, and 5-hydroxymethyluridine. m5C and m5U did not change the fidelity of T7 RNA polymerase in ProtoScript II. However, m6A, pseudouridine, and hm5U increase the first strand error rates. In addition to measuring these error rates, we're also able to identify the types of substitution errors introduced by these modifications.
For a more detailed explanation of the types of errors introduced by these modifications, check out our recent paper published in Nucleic Acids Research.
To view our open access publication, check out the link in the post.
Related Videos
-
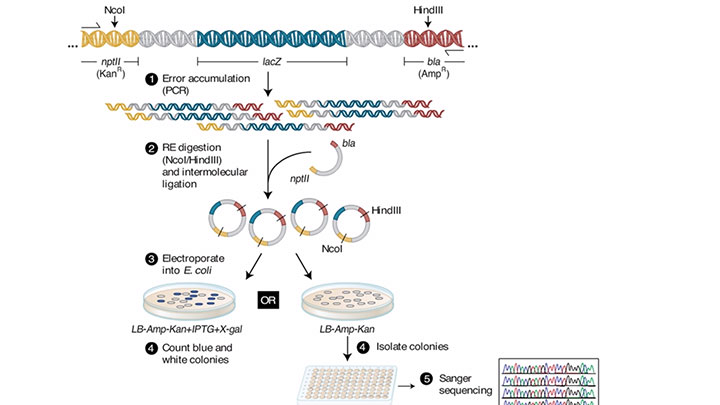
Fidelity testing workflow -

DNA Ligase Fidelity: When does it matter? -

Why Choose Q5® High-fidelity Polymerase?

