Guidelines for using NEBuilder® HiFi DNA Assembly
NEBuilder HiFi DNA Assembly is a robust and powerful tool that can be used to combine different pieces of DNA. This chart provides recommendations, such as ratios of insert to vector as well as incubation times, for various scenarios. These guidelines may need to be adjusted to suit your particular situation.
|
Inserts |
Vector |
Recommended Ratio (insert : vector)
Amount setup in 20 µl (insert : vector (in fmol)) |
Recommended Time for Assembly | Cloning Application Recommendations |
|---|---|---|---|---|
3 kb insert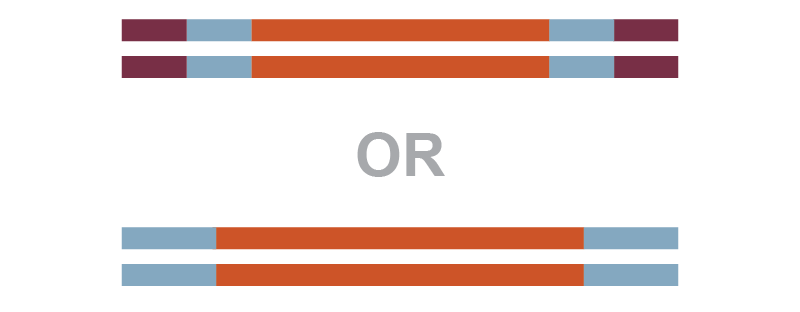
|
Linear 2.1 kb plasmid | 2 : 1 20 : 10 |
15-60 min |
|
| 120 bp < insert < 200 bp |
Linear 5.4 kb plasmid |
10-5 : 1 200-100 : 20 |
15-60 min |
|
|
750 bp x 4, 20-30 bp overlap |
Circular assembly |
1 : 1 : 1 : 1 20 : 20 : 20 : 20 |
15-60 min |
|
750 bp x 4, 15 bp overlap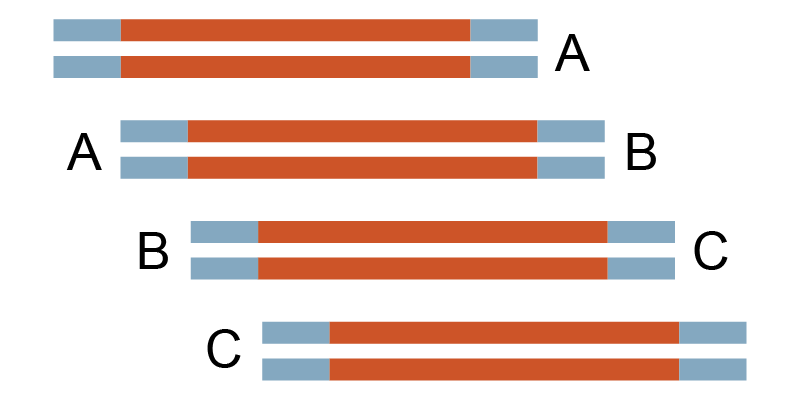 |
Circular assembly |
1 : 1 : 1 : 1 40 : 40 : 40 : 40 |
15-60 min |
|
1,000 bp x 3, 25 bp overlap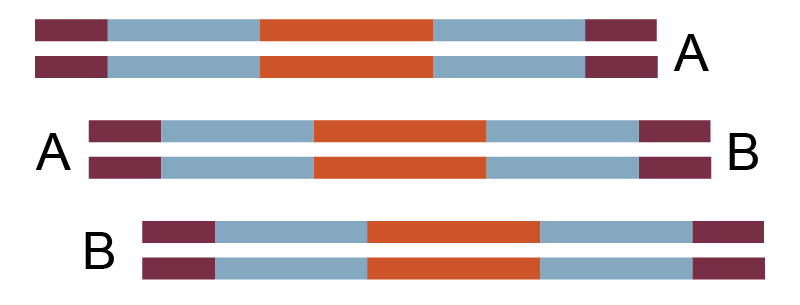 |
Linear 3.3 kb plasmid |
1 : 1 : 1 : 1 50 : 50 : 50 : 50 |
60 min |
|
6 kb x 2 insert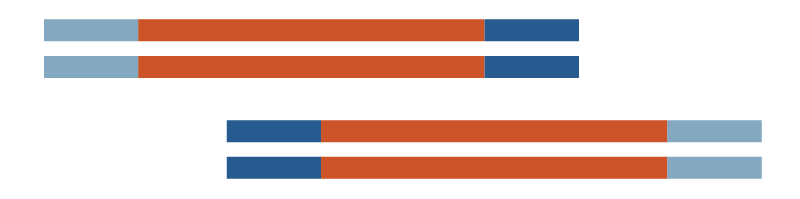 |
Linear 7 kb plasmid |
1 : 1 : 1 25 : 25 : 25 |
60 min |
|
| 70-mer ssDNA |
Linear 9.8 kb plasmid | 200-50 : 1 1000-500 : 10 |
15-60 min |
|
| 60-mer x 2 |
Linear 2.6 kb plasmid |
40-20 (annealed oligos) : 1 1000-500 : 25 |
15-60 min |
|
| 60-mer x 5 |
Linear 2.6 kb plasmid |
40-20 (annealed oligos) : 1 1000-500 : 25 |
15-60 min |
|
1000 bp x 5, 80 bp overlap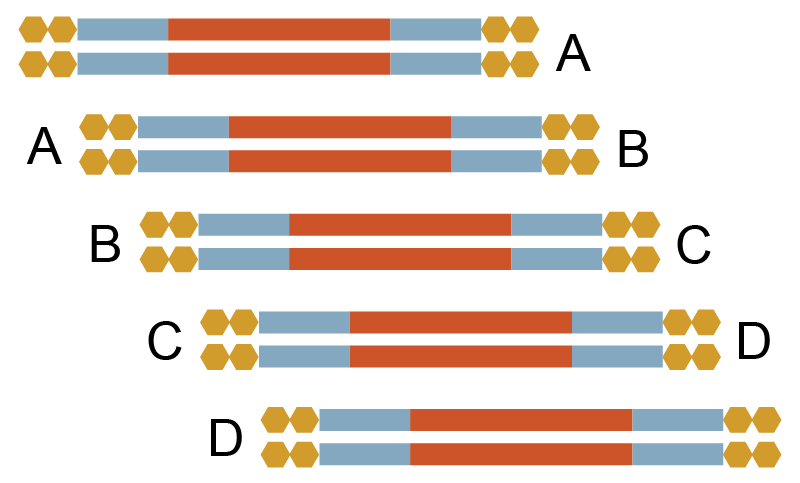 |
Linear 3.3 kb plasmid | 1 : 1 : 1 : 1 : 1 : 1 50 : 50 : 50 : 50 : 50 : 50 |
60 min |
|
450 bp x 11, 25 bp overlap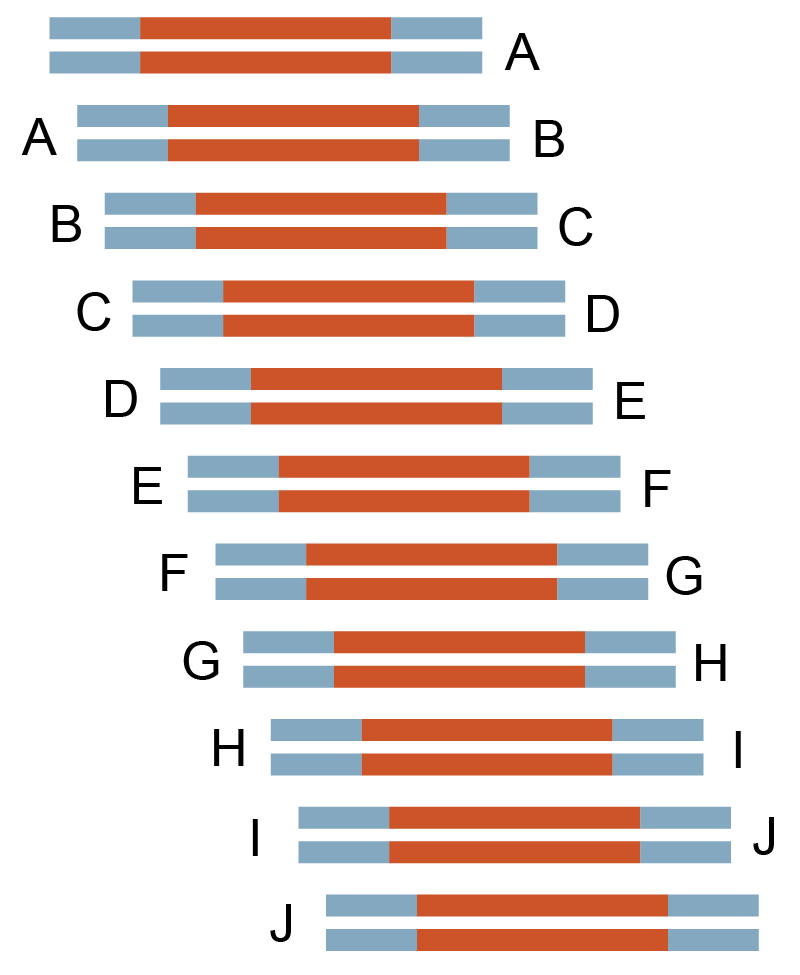 |
Linear 3.3 kb plasmid |
1 : 1 : 1 : 1 : 1 : 1 : 1 : 1 : 1 : 1 : 1 : 1 50 (each insert) : 50 |
60 min |
|