mRNA Decapping
Eukaryotic mRNAs are modified at their 5′ end early during transcription by the addition of N7- methyl guanosine (m7G), which forms the “cap” on the first 5′ nucleotide. Identification of the 5′ nucleotide on mRNA is necessary for determination of the transcription start site (TSS). We explored the effect of various reaction conditions on the activity of the yeast scavenger mRNA decapping enzyme DcpS and examined decapping of 30 chemically distinct cap structures varying the state of methylation, sugar, phosphate linkage, and base composition on 25mer RNA oligonucleotides. Contrary to the generally accepted belief that DcpS enzymes only decap short oligonucleotides, we found that the yeast scavenger decapping enzyme decaps RNA transcripts as long as 1400 nucleotides. Further, we validated the application of yDcpS for enriching capped RNA using a strategy of specifically tagging the 5′ end of capped RNA by first decapping and then recapping it with an affinity-tagged guanosine nucleotide.
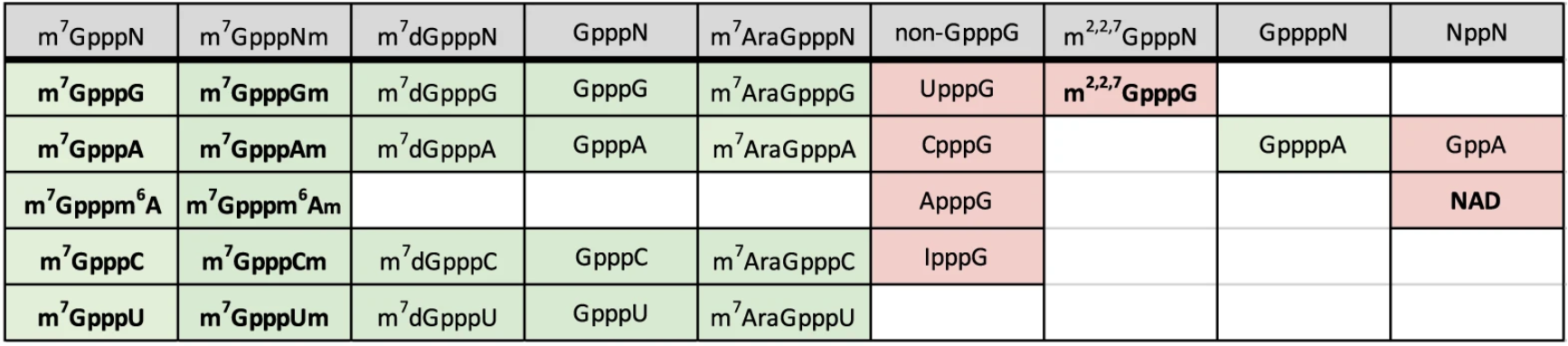
Decapping properties of yDcpS. Synthetic cap analog substrates efficiently decapped are shaded in green and substrates resistant to decapping (less than 10%) are shaded in pink. Substrates in boldface characters indicate canonical cap structures. Figure from Wulf et al. 2019.

