T7 DNA Ligase
T7 DNA Ligase will ligate these substrates:
dsDNA
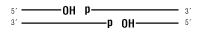
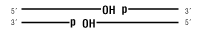
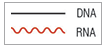
Nicked DNA/RNA
T7 DNA Ligase is an ATP-dependent dsDNA ligase from bacteriophage T7.
- Ligates sticky-ends only
- This product can be purchased in larger volumes. Submit an inquiry to find out more.
- Not sure which ligase to choose? Refer to our DNA and RNA Ligase Properties Chart or DNA Ligase Selection Chart
Featured Video
-
Product Information
T7 DNA Ligase is an ATP-dependent ds DNA ligase from bacteriophage T7. It will catalyze the formation of a phosphodiester bond between adjacent 5´ phosphate and 3´ hydroxyl groups of duplex DNA. Cohesive end ligation and nick sealing can be efficiently catalyzed by T7 DNA Ligase (1,2). However, unlike T4 and T3 DNA Ligases, blunt end ligation is not efficiently catalyzed by T7 DNA Ligase. Addition of high concentrations of PEG 6000 [≥ 20% (w/v)] to the reaction can force T7 DNA Ligase to have measurable activity. However, under typical reaction conditions blunt-end DNA ligation does not occur in the presence of T7 DNA Ligase, making it a good choice for applications in which blunt and cohesive ends of DNA are present but only the cohesive ends are to be joined.
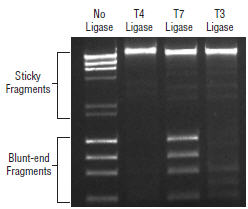
T7 DNA Ligase does not ligate blunt-end DNA fragments. Ligation reactions containing blunt fragments (ΦX174 DNA-HaeIII Digest (NEB #N3026 ) and sticky-end fragments (λ DNA-HindIII Digest (NEB #N3012 ) were set up with 200 ng of each substrate and 1 μl of each ligase, and incubated for 30 minutes at 25°C in their corresponding reaction buffers. Reactions were immediately stopped with 6X loading dye and resolved by electrophoresis on a 1% agarose gel and stained with ethidium bromide. Product Source
An E. coli strain containing a recombinant gene encoding T7 DNA Ligase.- This product is related to the following categories:
- DNA Ligases Products
- This product can be used in the following applications:
- Cloning Ligation,
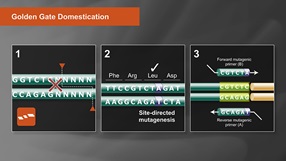
- NEBridge® Golden Gate Assembly
-
Protocols, Manuals & Usage
-
Tools & Resources
-
FAQs & Troubleshooting
-
Citations & Technical Literature
-
Quality, Safety & Legal
Other Products You May Be Interested In
Ineligible item added to cart
Based on your Freezer Program type, you are trying to add a product to your cart that is either not allowed or not allowed with the existing contents of your cart. Please review and update your order accordingly If you have any questions, please contact Customer Service at freezers@neb.com or 1-800-632-5227 x 8.