Gibson Assembly® Master Mix
Gibson Assembly Master Mix (NEB #M5510) has been reformulated with components containing Recombinant Albumin (rAlbumin) beginning with Lot #10229799. Learn more.
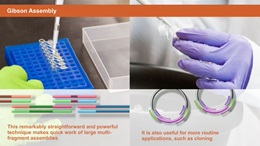
Have you tried NEBuilder HiFi DNA Assembly? NEBuilder HiFi offers several advantages over NEB Gibson Assembly. For more information, visit NEBuilderHiFi.com.Gibson Assembly allows for successful assembly of multiple DNA fragments, regardless of fragment length or end compatibility.
- Assembly and transformation in just under two hours
- Flexible sequence design (scar-less cloning)
- No PCR clean-up step required
- High transformation efficiencies for inserts up to 20 kb
- Easily adapted for multiple DNA manipulations, including site-directed mutagenesis
Featured Video
-
Product Information
Gibson Assembly was developed by Dr. Daniel Gibson and his colleagues at the J. Craig Venter Institute and licensed to NEB by Synthetic Genomics, Inc. It allows for successful assembly of multiple DNA fragments, regardless of fragment length or end compatibility. It has been rapidly adopted by the synthetic biology community due to its ease-of-use, flexibility and suitability for large DNA constructs.
Gibson Assembly efficiently joins multiple overlapping DNA fragments in a single-tube isothermal reaction (1,2). The Gibson Assembly Master Mix includes three different enzymatic activities that perform in a single buffer:- The exonuclease creates single-stranded 3´ overhangs that facilitate the annealing of fragments that share complementarity at one end (overlap region).
- The polymerase fills in gaps within each annealed fragment.
- The DNA ligase seals nicks in the assembled DNA.
To help select the best DNA assembly method for your needs, please use our Synthetic Biology/DNA Assembly Selection Chart.
For help designing primers, please view our primer design video.
Overview of the Gibson Assembly Method
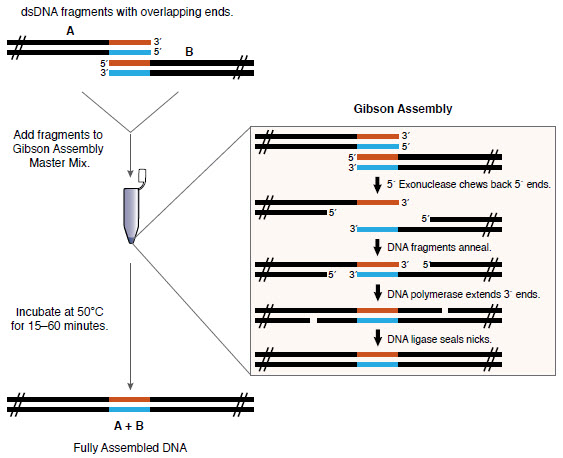
Specification:
10 μl of 2X Gibson Assembly Master Mix was incubated with 6 fragments (5 fragments of 400 bp and one of 2,780 bp, with 40 bp overlap, 0.05 pmol each) in a final volume of 20 μl at 50°C for 60 minutes. NEB 5-alpha Competent E. coli (NEB #C2987) were transformed with 2 μl of the master mix/fragment mixture using the transformation protocol. Greater than 100 white colonies were observed when 1/10 of the outgrowth was spread on an ampicillin plate with IPTG/Xgal and incubated overnight.
Overview of Gibson Assembly Master Mix Protocol:
- Design primers to amplify fragments (and/or vector) with appropriate overlaps
- PCR amplify fragments using a high-fidelity DNA polymerase.
- Prepare linearized vector by PCR amplification using a high-fidelity DNA polymerase or by restriction digestion.
- Confirm and determine concentration of fragments using agarose gel electrophoresis, a Nanodrop™ instrument or other method
- Add DNAs to Gibson Assembly Master Mix and incubate at 50°C for 15 minutes to 1 hour, depending on number of fragments being assembled.
- Transform into E. coli or use directly in other applications
- This product is related to the following categories:
- DNA Assembly, Cloning and Mutagenesis Kits Products
- This product can be used in the following applications:
- Gibson Assembly®
-
Protocols, Manuals & Usage
-
Tools & Resources
-
FAQs & Troubleshooting
-
Citations & Technical Literature
-
Quality, Safety & Legal
Featured Videos
-
Introduction to Gibson Assembly®
-

Gibson Assembly Workflow
-

NEBUILDER® Assembly Tool 2.0 What’s New?
-

NEBUILDER® Assembly Tool 2.0 Fragments Amplified by PCR
-

NEBUILDER® Assembly Tool 2.0 Restriction Enzyme Digest
-
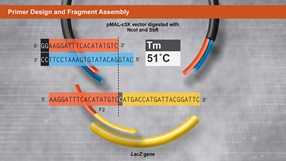
Primer Design and Fragment Assembly Using NEBuilder HiFi DNA Assembly® or Gibson Assembly®
Other Products You May Be Interested In
Ineligible item added to cart
Based on your Freezer Program type, you are trying to add a product to your cart that is either not allowed or not allowed with the existing contents of your cart. Please review and update your order accordingly If you have any questions, please contact Customer Service at freezers@neb.com or 1-800-632-5227 x 8.