RT-qPCR strategies for molecular diagnostics, OEM and customized solutions
Posted on Thursday, June 24, 2021
By
Topic: Tips for the lab
Scientists have quickly adapted amplification technologies to meet molecular diagnostic needs during COVID-19. Life science as a whole is in an epic endeavor to provide efficient and reliable solutions for developing diagnostic assays against the SARS-CoV-2 virus. According to the CDC COVID Data Tracker, over 440 million COVID-19 nucleic acid amplification tests have been performed in total in the United States alone. Really though, it's globally- scaled testing that will help prevent outbreaks and monitor vaccine effectiveness against SARS variants of concern. Since COVID infections can often be asymptomatic, and vaccinations are not yet universal, testing capacity is still urgently needed. From public health monitoring to medical care decision making – the technical aspects of optimizing nucleic acid amplification and detection of viral RNA feed directly into success with PCR-based MDx tests.
New England Biolabs Scientist Greg Patton, Ph.D., has had an important role in supporting SARS-CoV-2 RT-PCR / qPCR test developers using NEB’s Luna® amplification and detection reagents. I spoke with him recently on challenges in the field of PCR- based molecular diagnostics.

What are some of the latest research goals to improve RT-PCR-based molecular diagnostics for COVID-19?
One of the most important molecular diagnostics challenges is to create simpler workflows. Simplicity from start to finish helps to reduce test turnaround time – the time from patient sample collection to getting the results. This includes developing tests that can be run on commonly available equipment instead of specialized equipment. We can't really gain much traction on mass testing if your test requires an instrument that no one has. Simplified assays can also reduce reliance on consumables. The FDA put out last year that they would prioritize granting Emergency Use Authorization (EUA) to tests that were more widely deployable for good reason. Assay simplicity is the key to increasing massive testing capacity.
Another goal is accuracy….
Can you share a specific example of qPCR molecular diagnostics simplification?
Definitely. The field has seen a lot of tests move away from swab-based extraction workflows to simplified saliva-based protocols. Early in the pandemic, tests utilized a nasal pharyngeal swab, which is a deep nasal swab. If you want people to go and get tested multiple times, a painful sample collection process that’s like a brain scramble every time isn’t going to help. One of the common steps after sample collection is RNA extraction and purification. Simplified saliva protocols eliminate that step. Saliva samples can be incubated with Proteinase K and quickly heat-treated to inactivate the virus. Then that sample can go directly into the RT-qPCR workflow without the need for extraction and purification. It saves time and a decent amount of money. What’s more – it reduces your reliance on the associated reagents and extraction instruments. Saliva sample assays are simpler.
Saliva Direct is an EUA protocol from Yale University that uses NEB’s Luna reagents. It’s a unique EUA in that it’s a protocol and they have ultimately provided designations to other labs to run it. It saves a significant amount of time. Another great example is a multiplex RT-PCR for three different targets developed by the Lab at the University of California, Berkeley that also uses Luna reagents, called the IGI-LuNER. This workflow targets N, E, and RNase P as an internal control in a 384- well format using saliva samples.
How do researchers improve PCR test accuracy and eliminate false results?
First, to take a step back, no test is 100% accurate. Every test is going to have some rate of false positive and/or false negative results. It’s really about designing a robust test to minimize those false results. There are many factors to consider: sample type, stage of infection and the RT-qPCR workflow. Each assay is going to have an optimal design based on unique factors.
Noted, so what about false negatives in PCR testing?
In terms of false negatives, primer and probe selection for your target is really important. The key is designing an assay that’s highly specific and has a low limit of detection (LOD). LOD is a common term in molecular diagnostics. It's the lowest amount of target that can be detected. And often, it's defined at 95% confidence. If you have a hundred positive patient samples at the LOD, you will pick up at least 95% of them. The theoretical limit of detection for RT-qPCR or nucleic acid technologies in general is three copies. Getting down to single digit copy detection is very helpful to minimize any sort of false negative results. An assay with an LOD of one copy is going to be much more sensitive than say, one thousand copies. That’s why each set of primer-probe designs for nucleic acid amplification technology is critical.
What causes false positives?
In molecular diagnostics testing for SARS-CoV-2, false positives are often about lab error and cross-contamination. The main source in PCR diagnostics of cross-contamination is amplicon contamination. You can think of molecular amplification like running an office copier machine. As if you are taking one sheet of paper and copying it over, and over, and over again. As you do that, you run the risk of those copies ultimately spreading in the laboratory through aerosolization or whatnot. Suddenly a future reaction or sample gives you false positive results.
How can MDx laboratories prevent cross-contamination?
Often, molecular diagnostics labs are all set up in a united, directional workflow. Sample processes and reagents are in one area. The actual PCR amplification is done in another location. Analysis in yet another space. You want to contain activities to completely separate locations. The benefit of real-time PCR is that amplification and analysis all take place in a closed system. You never have to open the tubes for analysis. Opening the tubes is the primary source of contamination – when the amplicon could be accidently aerosolized, traveling into the lab space.
NEB reagents utilizes a solution to what’s commonly referred to as carryover contamination prevention. It’s the incorporation of dUTP or uracil, and Thermolabile uracil DNA glycosylase (UDG) into the actual amplification reaction. That way, when you amplify a target, that amplicon will then contain uracil. What that means is that if that amplicon were to escape and actually contaminate a future PCR reaction, UDG will recognize uracil and remove it. Once the uracil has been removed from the amplicon, that amplicon is no longer amplifiable. That minimizes false positives.
Is there a possibility that PCR reagents themselves can be a source of contamination?
Yes, this is possible. Contamination of a component was likely the cause of failure of the early COVID-19 testing kits made by the CDC. We are scrupulous in avoiding cross-contamination. Our reagents have been adopted into so many SARS-CoV-2 testing workflows, quality control takes on a humanitarian urgency.
What are the strengths of qPCR multiplexing for molecular diagnostics?
Multiplexing is great because you get more answers for a single sample input. For example, let’s say a patient presents in the middle of flu season with symptoms that could fit both the flu and SARS-CoV-2. It’s nice to interrogate a single sample in multiple ways to see if the patient has one virus or the other, or both. Multiplexing helps conserve a precious sample by running one test instead of two (one for flu, one for SAR-CoV-2).
Multiplexing can also increase testing capacity. For example, the Luna SARS-CoV-2 RT-qPCR Multiplex Assay Kit is based on the CDC assay for SARS-CoV-2 detection and looks at three targets; N1, N2 and human RNase P in a single reaction. In the absence of this multiplexed reaction, a 96-well plate can look at only 32 samples for all three targets in that plate. Multiplexing allows you to run up to 96 patient samples in a 96-well plate. The outcome is that you get more results per sample and reduced assay costs because you are testing more samples overall using fewer PCR reagents.
What are the weaknesses of multiplexing?
The negative aspect of multiplexing is that it requires a decent amount of optimization. Because there are multiple primers and probes together in the same reaction, you must ensure they are compatible. They need to work well in the context of all reactions occurring in a single tube. One problem that can come up is that one of the targets dominates the reaction. You need to carefully balance the concentration of all primers and probes in addition to the sequence context so that you get robust amplification of all targets in the same tube. Thus, multiplexing requires a lot more optimization upfront.
Why are the Luna reagents so useful for qPCR multiplexing?
One of the strengths of NEB’s Luna RT-qPCR reagents is that they can do 5-plex reactions such that a single sample can be integrated in five different ways. Up until recently, 5-plex was the limit because real-time PCR instruments typically have five fluorescent channels and one target is detection in each channel. The instruments are often limited to five channels to get good spectral separation of the excitation and emission of the fluorophores used for detection. However, scientists are starting to push the limits of the instrumentation and coming up with creative ways to detect multiple targets per channel. If you can do it molecularly, you don’t want to be held back by the engineering.
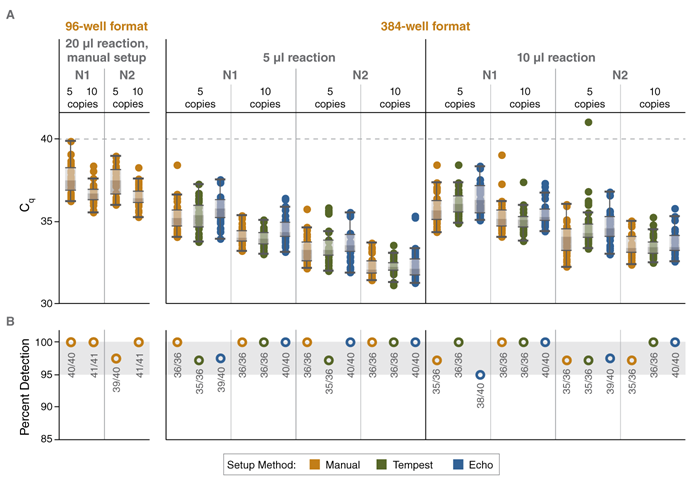
FIGURE: Automation-compatible, high-throughput multiplex RT-qPCR for detection of SARS-CoV-2 RNA with high sensitivity using Luna reagents
Cq values (A) and percent detection (B) for multiplex SARS-CoV-2 assays assembled in 96-well or 384-well format either manually or using the
indicated automated liquid handling platform. Reactions contained either 5 or 10 copies of synthetic SARS-CoV-2 RNA. An LOD of 5 copies/
reaction was maintained for all assembly workflows. Data shown were collected using Bio-Rad CFX96 and CFX384 real-time instruments.
That’s a good point. So, exactly what are the unique technical advantages of Luna amplification and detection?
Aptamer-based enzyme control is a technology that shuts down polymerase activity while assays are being set up at room temperature, which is very handy for set up and especially automation testing protocols. It's basically a small nucleic acid that binds to the polymerase and shuts it down at room temperature. Then, as you load the assays into the thermocycler, the heat causes the aptamer to disassociate from the polymerase. This flexibility can prevent problems in commercial testing in a clinical lab setting.
I already mentioned that Luna is compatible with carryover prevention. The reagents also contain a universal passive reference dye that ensures compatibility on a variety of real-time instruments.
There continues to be pandemic-related lab supply chain issues. How are RT-qPCR test developers approaching that issue?
From a science perspective, this is really the same theme of assay accessibility and scale-up. Ideally, simpler workflows will rely on fewer consumables. Again, protocols that skip RNA extraction reduce consumables upfront. Multiplexing can run more samples with fewer lab consumables. At NEB, we produced an application note for using the Beckman acoustic handler – that conserves materials by assay miniaturization. Acoustic handler instruments are also amazing because they eliminate pipette tips. I never thought our field would be in a position where it would be difficult to get 96-well PCR plates or pipette tips, but this happened in some places during the pandemic. From our own perspective, NEB had the capabilities to ramp up reagent supplies to meet pandemic needs for customers without any conflict of supplying our own. That’s been reassuring for developers from the start.
Detection of the virus and assay technical performance, design, and validation, is just one piece of the overall puzzle for testing.
Is there any aspect to RT-PCR virus detection that molecular diagnostics developers don't often consider upfront?
Detection of the virus and assay technical performance, design, and validation, is just one piece of the overall puzzle for testing. We’ve seen quite frequently that developers can be very focused on the performance of a test or a particular technology and they fail to consider some of the bigger picture items like manufacturing, distribution, implementation, or other challenges.
Some developers come to NEB late in their development phase, ready to execute only to discover that their chosen path might not be ideal for any number of reasons. For example, they incorporate a suboptimal product into their workflow from a technical perspective or one that is difficult to meet their forecasted demands on short notice. Often, we can direct them to the best product that would meet their needs immediately from both a manufacturing and technical perspective, but that’s always a difficult conversation to have late in the game. Instead, we recommend having conversations early with suppliers because it's much easier to guide folks on a path to success.
It sounds like there is more that NEB helps with than just test performance.
Exactly, we can make projects more efficient through collaboration from science to manufacturing.
NEB has a wealth of knowledge that we are always willing to share with the greater scientific community (and do so frequently through publications, our technical support system, NEB.com, and our customer interactions), so I would encourage developers to get in touch with us early on to leverage our expertise.
How do SARS-CoV-2 variants affect primer-probe designs?
This is an important consideration in assay robustness. There are two ways we need to look at coronavirus genetic drift.
Let’s say there is a mutation in the probe region, or the 3-prime end of your primer, your assay may fail completely if it only evaluates a single region of the SARS-CoV-2 genome, and the result will be a false negative. Test developers need to be mindful of coronavirus variants – to be certain that their primers and probes still work on any viral variants that pop up.
On the flip side, we also need assays to discern if you have a viral variant of concern. While this is an area of work that is heavily next generation sequencing-focused, developers continue to design PCR assays that will discriminate among several of the SARS-COV-2 mutant strains.
Beyond reagents, NEB now has a pretty powerful PCR Primer Monitor tool for MDx developers to monitor that their assays work for any SARS-CoV-2 mutations. The newly released White Paper Primer Monitor: an online tool to track SARS-CoV-2 variants that may impact primers used in diagnostic assays is a great guide on this new tool.
What has been the most rewarding part of your team’s work during the pandemic?
Knowing that we are assisting and helping in a humanitarian effort to get testing deployed. It gives people the chance to isolate themselves if they have COVID-19 and not pass it on to someone who is susceptible to the virus. To see the work that we have been doing translate into testing capacity and helping people in the real world right now… I’m thrilled. It’s amazing to think that the reagents we have worked on developing can benefit people. It’s meaningful to support test developers to help save lives.
NEB will not rent, sell or otherwise transfer your data to a third party for monetary consideration. See our Privacy Policy for details. View our Community Guidelines.
Products and content are covered by one or more patents, trademarks and/or copyrights owned or controlled by New England Biolabs, Inc (NEB). The use of trademark symbols does not necessarily indicate that the name is trademarked in the country where it is being read; it indicates where the content was originally developed. See www.neb.com/trademarks. The use of these products may require you to obtain additional third-party intellectual property rights for certain applications. For more information, please email busdev@neb.com.
Don’t miss out on our latest NEBinspired blog releases!
- Sign up to receive our e-newsletter
- Download your favorite feed reader and subscribe to our RSS feed
Be a part of NEBinspired! Submit your idea to have it featured in our blog.



