Luna® Universal qPCR & RT-qPCR
Experience best-in-class performance

All NEB products undergo rigorous testing to ensure optimal performance, and Luna is no exception. We took into consideration numerous important traits when evaluating qPCR, including specificity, sensitivity, accuracy and reproducibility, to develop best-in-class qPCR products. Furthermore, we did a comprehensive evaluation of many commercially-available qPCR and RT-qPCR products, and developed a method of analysis that allows you to quickly compare and evaluate the performance of these products. We wanted to be sure that Luna products will perform to your expectations for all of your targets.
Benefit from rigorous testing
- All Luna reagents were evaluated for specificity, sensitivity, efficiency, accuracy, linearity and reproducibility, and optimized for performance
- Development of novel "dots in boxes" analysis method for concise visual presentation of comprehensive evaluations
Use with ALL your targets
- Extensive panel testing and evaluation ensures reproducible results
- Luna reagents are optimized for strong performance across a wide range of targets and templates, including those of varied length, GC-content and abundance
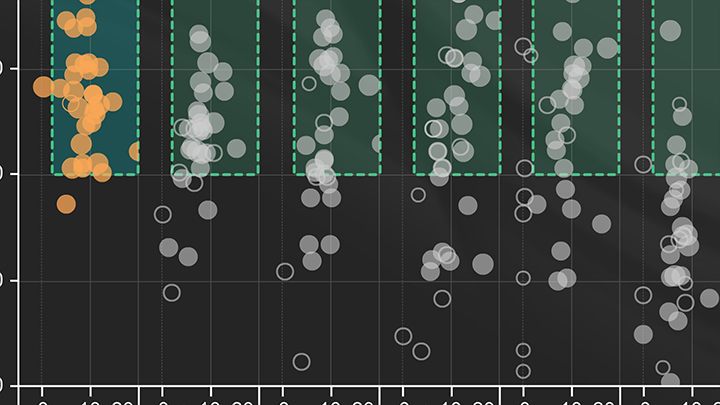
Evaluating qPCR results: capturing performance as "dots in boxes"
NEB has developed a method to better evaluate the large amount of qPCR data generated in an experiment. The output of this analysis is known as a dot plot, and captures the key features of a successful, high-quality qPCR experiment as a single point. This method of analysis allows many targets and conditions to be compared in a single graph.
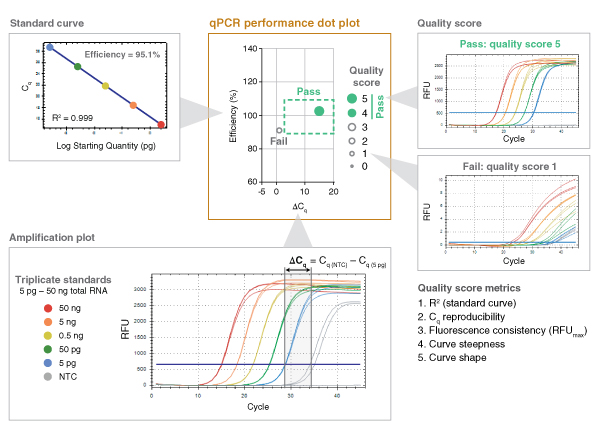
For each experiment, triplicate reactions are set up across a five-log range of input template concentrations (Amplification plot, bottom-left). Three non-template control (NTC) reactions are also included, for a total of 18 reactions per condition/target. Efficiency (%) is calculated (Standard curve, top-left) and is plotted against ΔCq (dot plot, center), which is the difference between the average Cq of the NTC and the lowest input. This parameter captures both detection of the lowest input and non-template amplification. Acceptable performance criteria are defined as an Efficiency of 90-110% and a ΔCq of ≥ 3 (green box).
Other performance criteria are captured using a 5-point Quality Score (top-right). Included are:
- Linearity of amplification, as indicated by the R2 standard curve
- Reproducibility, as indicated by the consistency of triplicate Cq values for each input concentration
- Fluorescence consistency, as indicated by similar endpoint fluorescence (RFUmax)
- Curve steepness
- Sigmoid curve shape
Quality Score is represented by the size and fill of the plotted dot, with experiments that pass all performance criteria represented by a solid dot within the box. These scoring methods were built upon the MIQE qPCR/RT-qPCR guidelines (Bustin, S.A. et al. (2009) Clin Chem. 55, 611-22 and Trombley Hall, A. et al. (2013) PLoS One 8(9):e73845). An example of this analysis is shown below.
Breaking it down: how we translate qPCR data into "dots in boxes"
|
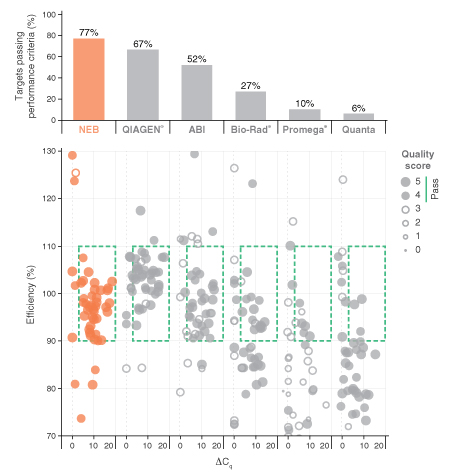
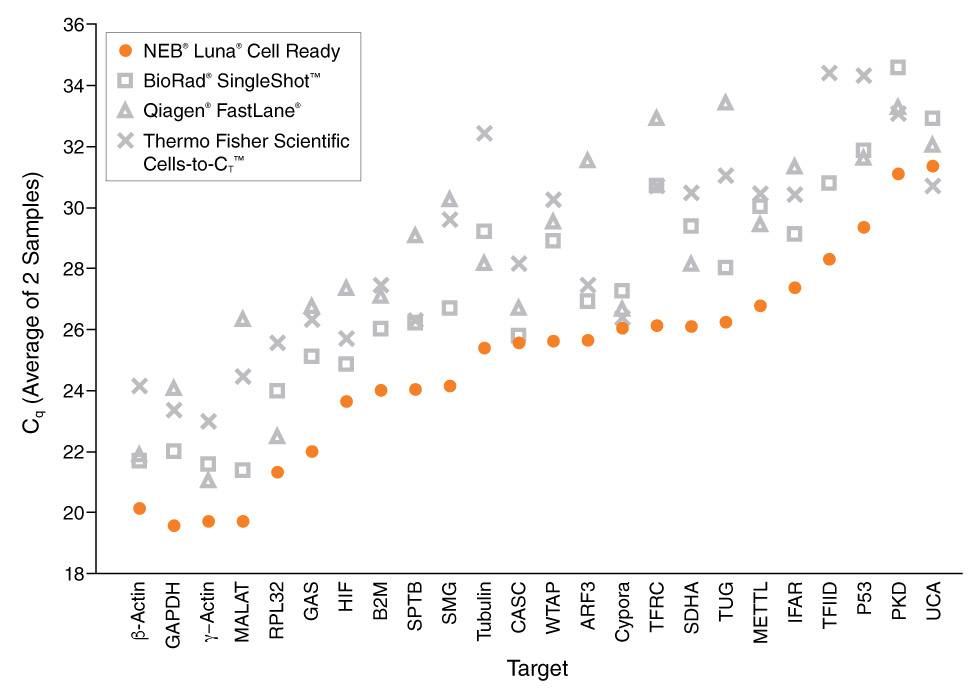
See how Luna products stack up
Comprehensive market-wide evaluation of qPCR and RT-qPCR products reveals that Luna outperforms the competition.
Available products:
| Luna Universal qPCR Master Mix Rapid, sensitive, and precise dye-based qPCR detection and quantitation of target DNA and cDNA sequences |
|
| Luna Universal Probe qPCR Master Mix Rapid, sensitive, and precise probe-based qPCR detection and quantitation of target DNA and cDNA sequences |
|
| Luna Universal One-Step RT-qPCR Kit Includes everything you need for rapid, sensitive, and precise dye-based qPCR detection and quantitation of RNA targets |
|
| Luna Universal Probe One-Step RT-qPCR Kit Includes everything you need for rapid, sensitive, and precise probe-based qPCR detection and quantitation of RNA targets |
|
| Luna Probe One-Step RT-qPCR 4X Mix with UDG 4X mix enables sensitive detection of target RNA sequences, with robust performance in multiplex applications of up to 5 targets |
|
| LyoPrime Luna Probe One-Step RT-qPCR Mix with UDG Lyophilized format enables sensitive detection of target RNA sequences with room temperature stability |
|
| Luna SARS-CoV-2 RT-qPCR Multiplex Assay Kit For real-time qualitative detection of SARS-CoV-2 nucleic acid using hydrolysis probes |
|
| LunaScript RT SuperMix Kit Single tube with blue tracking dye includes everything needed for first strand cDNA synthesis in two-step RT-qPCR workflows |
|
| LunaScript RT Master Mix Kit (Primer-free) Single-tube 5X master mix contains Luna Reverse Transcriptase, dNTPs, and Murine RNase Inhibitor |
|
| Luna Cell Ready One-Step RT-qPCR Kit All the necessary components for direct, dye-based RNA detection and quantitation from cell lysate, bypassing the need for RNA extraction and purification |
|
| Luna Cell Ready Probe One-Step RT-qPCR Kit All the necessary components for direct, probe-based RNA detection and quantitation from cell lysate, bypassing the need for RNA extraction and purification |
|
| Luna Cell Ready Lysis Module Coordinates cell lysis, RNA release, and genomic DNA removal simultaneously, resulting in cell lysate ready for direct RT-qPCR analysis with Luna Universal One-Step RT-qPCR kits |